Research teams
Molecular bases of human diseases
Machine learning and gene regulation

High-throughput sequencing experiments generate vast amounts of data, yet most analysis pipelines reduce this richness to simple gene counts aligned against a reference genome — discarding the very signals that make each patient, each cell, and each disease unique. Our lab takes a different approach. We work directly with raw sequence data, using k-mer decomposition and advanced machine learning to discover biological patterns that reference-based methods miss: rare variants, unannotated splicing events, non-coding signals, and patient-specific regulatory signatures.
This philosophy has led us to build widely-used tools such as iMOKA, which uses adaptive entropy filtering and coloured De Bruijn graphs to classify large sequencing cohorts, and IRFinder-S, the international reference software for detecting intron retention — a form of gene regulation we helped establish as a major post-transcriptional mechanism in cell differentiation and disease. More recently, we have shown that adversarial autoencoders can generate realistic synthetic human genomes, opening new avenues for privacy-preserving genomic research.
We are now pushing into the era of foundation models and single-cell AI. Our current projects tackle some of the most exciting questions at the interface of computation and biology: How can we predict the impact of a genetic variant on splicing in a specific cell type, when current tools are completely blind to cellular context? Can we integrate transcriptomic, epigenomic, and proteomic data from thousands of patients through a single AI framework to uncover the molecular determinants of rare phenotypes like spontaneous HIV control? And what can long-read sequencing from Oxford Nanopore reveal about previously invisible modes of intron retention — including coordinated clusters of retained introns with distinct RNA methylation and chromatin signatures?
We are a computational team embedded within a world-class genetics institute, giving us direct access to biological expertise, clinical collaborations, and cutting-edge sequencing data. We collaborate with algorithmicists, virologists, oncologists, and immunologists across France, Europe, and Australia. If you are excited about building AI methods that make real biological discoveries — and want to work at the frontier where deep learning meets gene regulation — we’d love to hear from you.
Alban Mancheron
Standard genomic analyses begin by aligning reads to a reference genome — a step that discards rare variants, unannotated isoforms, and any sequence not already in the reference. We bypass this entirely by decomposing sequencing data directly into k-mers, short overlapping words extracted from raw reads, and feeding them into machine learning models that learn which sequences matter. Our tools GECKO and iMOKA use genetic algorithms, adaptive entropy filters, and coloured De Bruijn graphs to sift through billions of k-mers and pinpoint the handful that distinguish disease from healthy, responder from non-responder. Because our approach is agnostic to the reference, it captures signals that conventional pipelines cannot see — from cryptic splice variants to pathogen-derived sequences integrated into the host genome.
Intron retention as a regulatory mechanism
Andrew Oldfield
Intron retention occurs when an intron remains in the mature mRNA instead of being spliced out. Once dismissed as noise, it is now recognised as a widespread and finely tuned layer of gene regulation. Our work helped put intron retention on the map, from demonstrating its orchestrated role in blood cell differentiation to uncovering how DNA methylation and MeCP2 control which introns are retained. We develop the IRFinder suite, the most widely used software for detecting and quantifying intron retention from short-read and long-read RNA-seq data. Our latest long-read analyses using Oxford Nanopore are revealing something unexpected: introns retained alone and introns retained in coordinated clusters within the same transcript appear to be two fundamentally different phenomena, with distinct RNA methylation profiles and chromatin signatures — suggesting separate regulatory mechanisms that we are now working to dissect.
Cell-type-aware prediction of gene regulation
A genetic variant can disrupt splicing in neurons while being completely harmless in liver cells, yet every existing predictor gives one answer regardless of cell type. We are building AI architectures that are aware of cellular identity. By combining genomic sequence encoders with cell embeddings from foundation models like Geneformer — transformers pre-trained on millions of single cells that implicitly capture which splicing factors and regulatory programs are active in each cell type — we aim to predict variant effects on splicing with cell-type resolution. Our first application targets autism-associated variants across cortical cell populations, but the framework is designed to be general: any regulatory process, any cell type, any disease.
Generating realistic artificial genomes
Callum Burnard
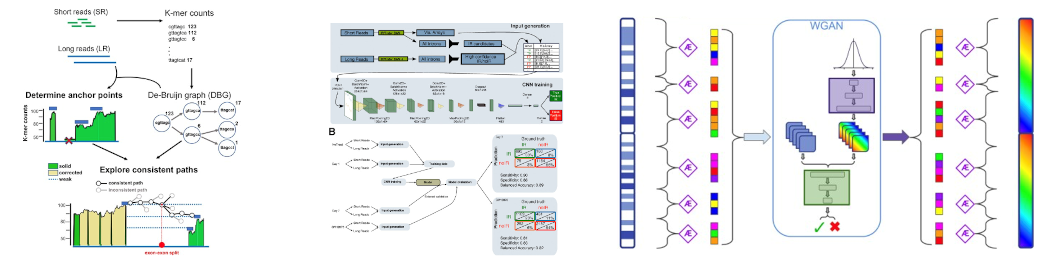
A published human genome is a scientific resource and a privacy risk — it can be used to re-identify donors and infer sensitive health information about their relatives. We address this by generating synthetic genomes that are scientifically useful but belong to no real person. Our approach segments the genome along natural recombination hotspots to preserve biologically meaningful haplotype blocks, encodes each block with optimised variational autoencoders, and generates novel genomes using adversarial networks. The resulting synthetic populations faithfully reproduce allele frequencies, linkage disequilibrium patterns, and population structure while being verifiably distinct from any real individual. We are now extending this framework to multi-omic data — synthetic genotype-transcriptome-methylome profiles — to enable secure data sharing in an era of increasing genomic privacy regulation.


- Burnard C, Mancheron A and Ritchie W (2025) Generating realistic artificial human genomes using adversarial autoencoders. NAR Genomics and Bioinformatics, 7(3), lqaf101
- Lorenzi C*, Barriere S*, Arnold K, Luco RF, Oldfield AJ and Ritchie W (2021) IRFinder-S: a comprehensive suite to discover and explore intron retention. Genome Biology, 22(1), 307
- Villemin JP, Lorenzi C, Cabrillac MS, Oldfield A, Ritchie W and Luco RF (2021) A cell-to-patient machine learning transfer approach uncovers novel basal-like breast cancer prognostic markers amongst alternative splice variants. BMC Biology, 19(1), 70
- Grabski DF*, Broseus L*, Kumari B*, Rekosh D, Hammarskjold ML and Ritchie W (2021) Intron retention and its impact on gene expression and protein diversity: A review and a practical guide. Wiley Interdisciplinary Reviews RNA, 12(1), e1631
- Augustus M, Pineau D, Aimond F, Azar S, Lecca D, Scamps F, Muxel S, Darlix A, Ritchie W, Gozé C, Rigau V, Duffau H and Hugnot JP (2021) Identification of CRYAB+ KCNN3+ SOX9+ Astrocyte-Like and EGFR+ PDGFRA+ OLIG1+ Oligodendrocyte-Like Tumoral Cells in Diffuse IDH1-Mutant Gliomas and Implication of NOTCH1 Signalling in Their Genesis. Cancers, 13(9), 2107
- Lorenzi C, Barriere S, Villemin JP, Dejardin Bretones L, Mancheron A and Ritchie W (2020) iMOKA: k-mer based software to analyze large collections of sequencing data. Genome Biology, 21(1), 261
- Broseus L, Thomas A, Oldfield AJ, Severac D, Dubois E and Ritchie W (2020) TALC: Transcript-level Aware Long Read Correction. Bioinformatics, 36(20), 5000–5006
- Broseus L and Ritchie W (2020) Challenges in detecting and quantifying intron retention from next generation sequencing data. Computational and Structural Biotechnology Journal, 18, 501–508
- Grasso G, Higuchi T, Mac V, Barbier J, Helsmoortel M, Lorenzi C, Sanchez G, Bello M, Ritchie W, Sakamoto S and Kiernan R (2020) NF90 modulates processing of a subset of human pri-miRNAs. Nucleic Acids Research, 48(12), 6874–6888
- Thomas A, Barriere S, Broseus L, Brooke J, Lorenzi C, Villemin JP, Beurier G, Sabatier R, Reynes C, Mancheron A and Ritchie W (2019) GECKO is a genetic algorithm to classify and explore high throughput sequencing data. Communications Biology, 2, 222
- Wong NKP, Cheung H, Solly EL, Vanags LZ, Ritchie W, Nicholls SJ, Ng MKC, Bursill CA and Tan JTM (2018) Exploring the Roles of CREBRF and TRIM2 in the Regulation of Angiogenesis by High-Density Lipoproteins. International Journal of Molecular Sciences, 19(7), 1903
- Rajasekhar M, Schmitz U, Broseus L, Wong JJL, Rasko JEJ, Ritchie W and Holst J (2018) Identifying microRNA determinants of human myelopoiesis. Scientific Reports, 8, 7264
- Barbier J, Chen X, Sanchez G, Cai M, Helsmoortel M, Higuchi T, Giraud P, Contreras X, Yuan G, Feng Z, Nait-Saidi R, Deas O, Bluy L, Judde JG, Rouquier S, Ritchie W, Sakamoto S, Xie D and Kiernan R (2018) An NF90/NF110-mediated feedback amplification loop regulates dicer expression and controls ovarian carcinoma progression. Cell Research, 28(5), 556–571
- Wong JJL, Gao D, Nguyen TV, Kwok CT, van Geldermalsen M, Middleton R, Pinello N, Thoeng A, Nagarajah R, Holst J, Ritchie W and Rasko JEJ (2017) Intron retention is regulated by altered MeCP2-mediated splicing factor recruitment. Nature Communications, 8, 15134
- Middleton R, Gao D, Thomas A, Singh B, Au A, Wong JJL, Bomane A, Cosson B, Eyras E, Rasko JEJ and Ritchie W (2017) IRFinder: assessing the impact of intron retention on mammalian gene expression. Genome Biology, 18(1), 51
- Schmitz U, Pinello N, Jia F, Alasmari S, Ritchie W, Keightley MC, Shini S, Lieschke GJ, Wong JJL and Rasko JEJ (2017) Intron retention enhances gene regulatory complexity in vertebrates. Genome Biology, 18(1), 216
- Sierro F, Evrard M, Rizzetto S, Melino M, Mitchell AJ, Florido M, Beattie L, Walters SB, Tay SS, Lu B, Holz LE, Roediger B, Wong YC, Warren A, Ritchie W, McGuffog C, Weninger W, Le Couteur DG, Ginhoux F, Britton WJ, Heath WR, Saunders BM, McCaughan GW, Luciani F, MacDonald KPA, Ng LG, Bowen DG and Bertolino P (2017) A Liver Capsular Network of Monocyte-Derived Macrophages Restricts Hepatic Dissemination of Intraperitoneal Bacteria by Neutrophil Recruitment. Immunity, 47(2), 374–388.e6
- Ritchie W (2017) microRNA Target Prediction. Methods in Molecular Biology, 1513, 193–200
- Wong JJL, Au AYM, Gao D, Pinello N, Kwok CT, Thoeng A, Lau KA, Gordon JEA, Schmitz U, Ritchie W, Holst J and Rasko JEJ (2016) RBM3 regulates temperature sensitive miR-142-5p and miR-143 (thermomiRs), which target immune genes and control fever. Nucleic Acids Research, 44(6), 2888–2897
- Edwards CR, Ritchie W, Wong JJL, Schmitz U, Middleton R, An X, Mohandas N, Rasko JEJ and Wong JJL (2016) A dynamic intron retention program in the mammalian megakaryocyte and erythrocyte lineages. Blood, 127(17), e24–e34
- Coudre C, Alani J, Ritchie W, Marsaud V, Sola B and Cahu J (2016) HIF-1α and rapamycin act as gerosuppressant in multiple myeloma cells upon genotoxic stress. Cell Cycle, 15(16), 2174–2182
- Li J, Teo AWJ, Goh KY, Tan JTM, Chan PML, Jiao H, Ritchie W, Rasko JEJ and Bursill CA (2016) The Poly-cistronic miR-23-27-24 Complexes Target Endothelial Cell Junctions: Differential Functional and Molecular Effects of miR-23a and miR-23b. Molecular Therapy Nucleic Acids, 5, e354
- van Geldermalsen M, Wang Q, Nagarajah R, Marshall AD, Thoeng A, Gao D, Ritchie W, Feng Y, Bailey CG, Dez-Peña N, Mañes S, deFazio A, Holst J and Rasko JEJ (2016) ASCT2/SLC1A5 controls glutamine uptake and tumour growth in triple-negative basal-like breast cancer. Oncogene, 35(24), 3201–3208
- Wong JJL, Au AYM, Ritchie W and Rasko JEJ (2016) Intron retention in mRNA: No longer nonsense: Known and putative roles of intron retention in normal and disease biology. BioEssays, 38(1), 41–49
- Wang Q, Hardie RA, Hoy AJ, van Geldermalsen M, Gao D, Fazli L, Sadeghi MC, Mees S, Gupta EK, Ritchie W, Guns EST, Cann B, Decker M, Hesse M, Holst J and Rasko JEJ (2015) Targeting ASCT2-mediated glutamine uptake blocks prostate cancer growth and tumour development. Journal of Pathology, 236(3), 278–289
- Kinjyo I, Qin J, Tan SY, Wellard CJ, Mrass P, Ritchie W, Doi A, Caber LL, Lladser A, Jackson JT, Herold MJ, Chopin M, Belz GT, Mempel TR, Weninger W and Hodgkin PD (2015) Real-time tracking of cell cycle progression during CD8+ effector and memory T-cell differentiation. Nature Communications, 6, 6301
* equal contribution





