Research teams
Genetics, cell biology and development department
RNA-protein dynamics in gene expression

Our lab studies the molecular mechanisms and dynamics of gene expression through RNA–protein interactions in living cells.
Gene expression is a highly dynamic and regulated process, in which RNAs and proteins assemble into dynamic macromolecular complexes across multiple spatial and temporal scales. These interactions determine how genetic information is translated into cellular states, while enabling cells to respond robustly to environmental changes. A central question of our research is how RNA-protein interactions are dynamically coordinated to control gene expression and maintain cellular homeostasis.
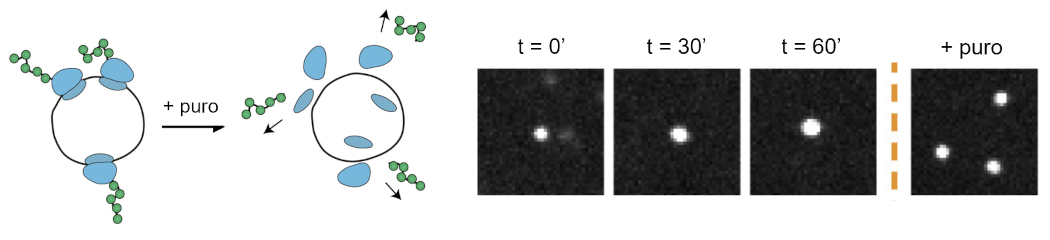

Real-time imaging of translation dynamics at the single-ribosome level in live U2OS cells. Adapted from Madern* and Yang* et al., Cell (2025).
To address this question, we combine quantitative biology with advanced single-molecule fluorescence imaging in living cells, allowing us to follow individual molecules in real time. We focus in particular on translation dynamics, ribosome collisions, and quality control pathways.
Capturing the transient and heterogeneous nature of RNA-protein interactions requires approaches that go beyond ensemble-averaged measurements. We develop new live-cell imaging methodologies that provide the resolution needed to dissect individual molecular interactions in their native cellular context, including nanotechnology-based approaches for single-molecule detection.
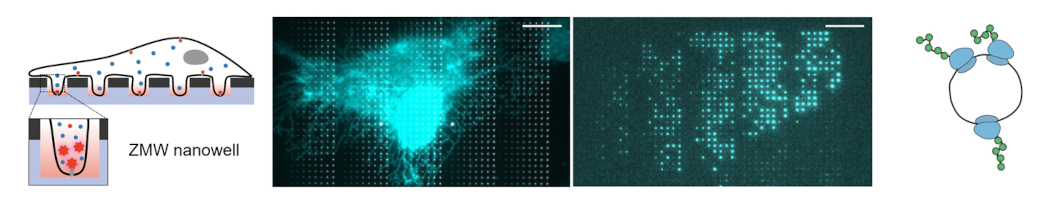
Live U2OS cell on the array of nanostructures called zero-mode waveguide (ZMW) nanowell for single-molecule fluorescence imaging. Adapted from Yang*, Klughammer*, and Barth* et al., ACS Nano (2023).
Ultimately, our research aims to uncover the molecular principles by which dynamic RNA–protein interactions encode regulatory information and ensure robust gene expression.




- Madern, M. F.†, Yang, S.†, Witteveen, O., Segeren, J., Bauer, M., and Tanenbaum, M. E. Long-term imaging of individual ribosomes reveals ribosome cooperativity in mRNA translation. Cell Volume 188, Issue 7, 1896 – 1911.e24 (2025).
- Tjeerdsma, R. B., Ng, T. F., Roorda, M., Bianchi, D., Yang, S., Bonnet, C., VanInsberghe, M., Everts, M., Bakker, F. J., de Boer, H. R., Moatti, N., Hustedt, N., Yin, J., Hoeg, L., Leibovitch, M., Sicheri, F., van Oudenaarden, A., de Jong, S., van den Berg, J., Tanenbaum, M. E., Brummelkamp, T. R., Durocher, D., and van Vugt, M. A. T. M. WEE1 inhibitors trigger GCN2-mediated activation of the integrated stress response. Nat Commun 16, 11598 (2025).
- Park, S.†, Yang, J.†, Yang, S.†, Han, Y. H.†, Kim, G., Seo, S. W., and Lee, N. K. Promoter-proximal transcription-translation coupling controls early transcription in Escherichia coli. Nucleic Acids Research Volume 53, Issue 17, gkaf896 (2025).
- Yi, S., Kim, E., Yang, S., Kim, G., Bae, D.-W., Son, S.-Y., Jeong, B.-G., Ji, J. S., Lee, H. H., Hahn, J.-S., Cha, S.-S., Yoon, Y. J., and Lee, N. K. Direct Quantification of Protein–Protein Interactions in Living Bacterial Cells. Adv. Sci. 12, 2414777 (2025).
- Son, J. B.†, Kim, S.†, Yang, S.†, Ahn, Y., and Lee, N. K. Analysis of Fluorescent Proteins for Observing Single Gene Locus in a Live and Fixed Escherichia coli Cell. J Phys Chem B 128(28), 6730-6741 (2024).
- Yang, S.†, Klughammer, N.†, Barth, A.†, Tanenbaum, M. E., and Dekker, C. Zero-mode waveguide nanowells for single-molecule detection in living cells. ACS Nano 17(20), 20179-20193 (2023).
- Yang, S.†, Kim, S.†, Kim, D. K., An, H. J., Son, J. B., Gynnå, A. H., and Lee, N. K. Transcription and translation contribute to gene locus relocation to the nucleoid periphery in E. coli. Nat Commun 10(1), 5131 (2019).
- Yang, S., Kim, S., Lim, Y. R., Kim, C., An, H. J., Kim, J. H., Sung, J., and Lee, N. K. Contribution of RNA polymerase concentration variation to protein expression noise. Nat Commun 5, 4761 (2014).




