Research teams
Molecular bases of human diseases
IMGT, international ImMunoGenetics information system®

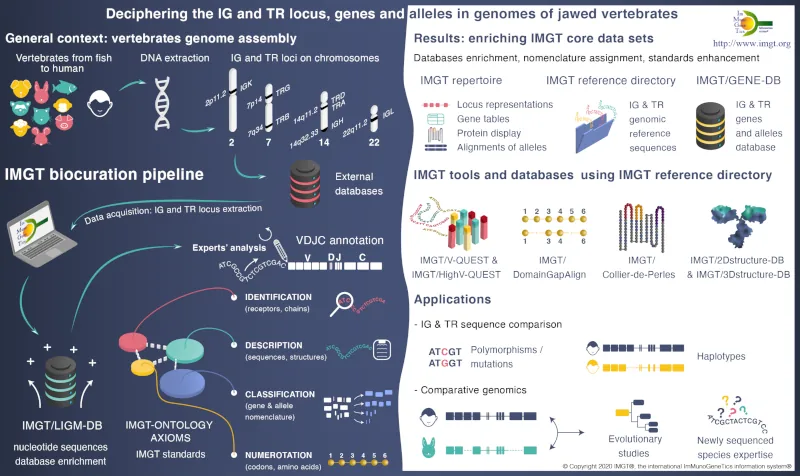
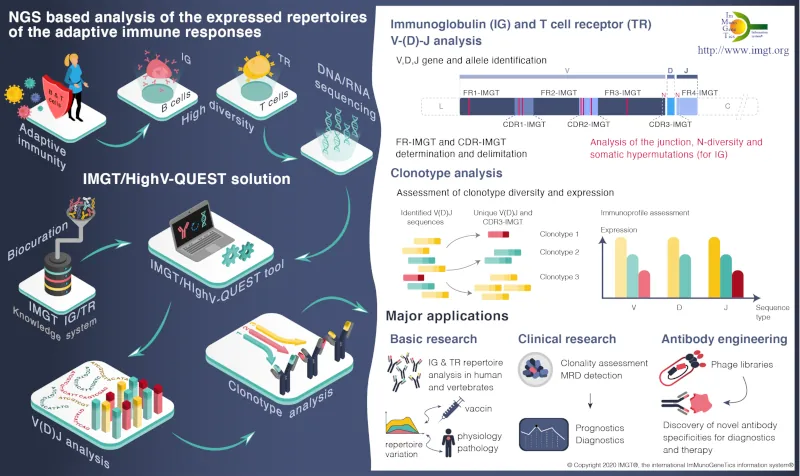
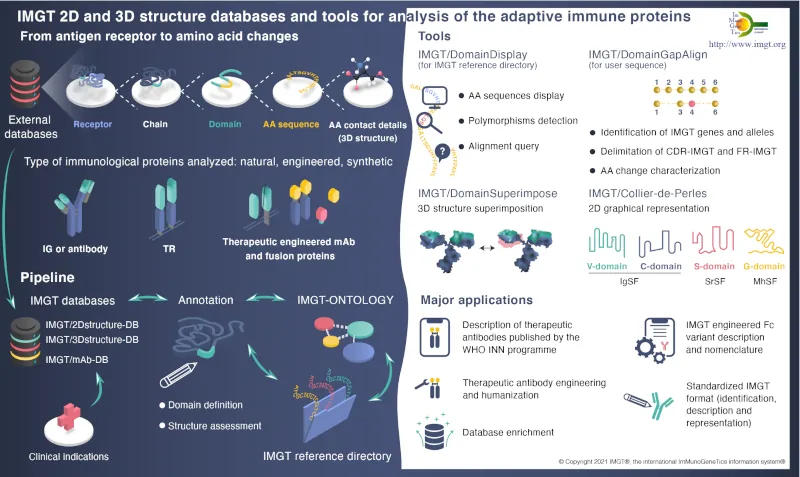
IMGT, is recognized as the mondial reference for the richness and quality of its expertized data. IMGT® is certified ISO 9001 and NF X50-900. The IMGT databases, online tools, and web resources of the information system are enriched by the results of the 3 research and development axes dedicated to the deciphering of antigen receptor genomic loci, the analysis of expressed repertoires, and the study of 2D and 3Dstructures of natural and engineered adaptive immune proteins.
it is widely used by academics and industrials involved in basic research (diversity and evolution of adaptive immune response in jawed vertebrates), in medical and veterinary research (analysis of antibody and T-cell receptor repertoires in normal and pathological situations), and for standardized description of properties of therapeutical monoclonal antibody.
Since July 1995, IMGT® is available on the Web. IMGT® is used by researchers and scientists, academic and from pharmaceutical and biotechnology companies, involved in fundamental research, medical research (autoimmune and infectious diseases, AIDS, leukemia, lymphoma, myeloma), veterinary research, genomics (genome diversity and evolution of the adaptive immune system), biotechnology related to antibody engineering for humanization of therapeutic antibodies, diagnostics (detection of minimal residual diseases) and therapeutic approaches (grafts, immunotherapy, vaccinology). The IMGT® Web server at Montpellier is accessed by more than 80,000 sites per year. IMGT® has an exceptional response with more than 150,000 requests per month.
Antibodies represent a large number of the pharmaceutical substances submitted to the World Health Organization International Nonproprietary Names (WHO INN) Programme. The INN definition of antibodies is based on the IMGT-ONTOLOGY concepts. Since 2008, amino acid sequences of monoclonal antibodies (mAb, INN suffix -mab), of fusion proteins for immune applications (FPIA) and composite proteins for clinical applications (CPCA) from WHO INN have been entered into IMGT®. These therapeutic applications emphasize the importance of the IMGT-ONTOLOGY concepts in bridging the gap between antibody sequences and 2D and 3D structures.
Another research interest, in collaboration mainly with the Unit of Medical Genetics, St-Joseph University, Beirut, and also with other teams in Tunisia and Algeria and the Children’s Hospital of Boston (Pr Raif GEHA) concerns rare Immuno Deficiencies and autosomal recessive genetic diseases in consanguineous families (there are as many as 25% of marriages between cousins, often first cousins and even double-first). The patients are autozygous (homozygous by descent) for very rare mutated genes and haplotypes, present in the common ancestor(s) of their parents. These exceptional genotypes are invaluable starting points to allow the identification more quickly of the yet unknown mutated genes. Their functions in the cell organization or in signaling pathways, including epigenetic and silencing by microARN, are unmasked and can be investigated. The genetic counselling can be performed in these families.
The better understanding of the molecular basis of the pathophysiology allows better choices in the development of diagnostic tools and innovative therapeutics. This research is beneficial not only for monogenic diseases, but also for complex ones. Indeed, the consanguinity, responsible also for homozygosity of large chromosomal regions, identical by descent, permits to discover more easily the genetic networks. This approach is also valid for the search of genetic susceptibility or protection against infectious diseases.




























































- Sanou G. , 1 Manso T., 1 Todorov K., 2 Giudicelli V., Duroux P. and Kossida S. Therapeutic Monoclonal Antibodies Repurposing in Oncology Domain using IMGT/mAb-KG Embeddings. Genomics, Proteomics & Bioinformatics 2025 (submitted).
- Kushwaha A., Duroux P., Giudicelli V., Todorov K., Kossida S. IMGT/RobustpMHC: robust training for class-I MHC peptide binding prediction. Briefings in Bioinformatics, 2024, Volume 25, Issue 6. doi: 10.1093.
- Debbagh C., Folch G., Jabado-Michaloud J., Giudicelli V., Kossida S. Deciphering Gorilla gorilla gorilla immunoglobulin loci in multiple genome assemblies and enrichment of IMGT resources. Front. Immunol., 2024, 15:1475003. doi: 10.3389/fimmu.2024.1475003.
- Sanou G., Manso T., Todorov K., Giudicelli V., Duroux P., Kossida S. IMGT/mAb-KG: the Knowledge Graph for Therapeutic Monoclonal Antibodies. Front. Immunol., 2024, 15, pp.1393839. doi: 10.3389/fimmu.2024.1393839.
- Golfinopoulou R., Giudicelli V., Manso T., Kossida S. Delving into Molecular Pathways: Analyzing the Mechanisms of Action of Monoclonal Antibodies Integrated in IMGT/mAb-DB for Myasthenia Gravis. Vaccines (Basel) 2023 Nov 26;11(12):1756. doi: 10.3390/vaccines11121756.
- Manso T., Kushwaha A., Nguefack Ngoune V., Georga M., Abdollahi N., Duroux P., Giudicelli V., Kossida S. Mechanisms of action of monoclonal antibodies in oncology integrated in IMGT/mAb-DB. Front. Immunol., 2023 May 05;14. doi:10.3389/fimmu.2023.1129323.
- Sanou G., Giudicelli V., Abdollahi N., Kossida S., Todorov K., Duroux P. IMGT-KG: A Knowledge Graph for Immunogenetics. The Semantic Web ISWC 2022. ISWC 2022. Lecture Notes in Computer Science, vol 13489. doi: 10.1007/978-3-031-19433-7_36.
- Nguefack Ngoune V., Bertignac M., Georga M., Papadaki A., Albani A., Folch G., Jabado-Michaloud J., Giudicelli V., Duroux P., Lefranc MP., Kossida S. IMGT® Biocuration and Analysis of the Rhesus Monkey IG Loci. Vaccines (Basel). 2022 Mar 3;10(3):394. doi: 10.3390/vaccines10030394 PMID: 35335026 Free PMC article.
- Manso T., Folch G., Giudicelli V., Jabado-Michaloud J., Kushwaha A., Nguefack Ngoune V., Georga M., Papadaki A., Debbagh C., Pégorier P., Bertignac M., Hadi-Saljoqi S., Chentli I., Cherouali K., Aouinti S., El Hamwi A., Albani A., Elazami Elhassani M., Viart B., Goret A., Tran A., Sanou G., Rollin M., Duroux P., Kossida S. IMGT® databases, related tools and web resources through three main axes of research and development. Nucleic Acids Res. 2022 Jan 7;50(D1):D1262-D1272. doi: 10.1093/nar/gkab1136 PMID: 34875068.
- Pégorier P., Bertignac M., Nguefack Ngoune V., Folch G., Jabado-Michaloud J.,Giudicelli V., Duroux P., Lefranc M.-P. and Kossida S. IMGT® Biocuration and Comparative Analysis of Bos taurus and Ovis aries TRA/TRD Loci. Genes (Basel), 2020a Dec 28;12(1):30. doi: 10.3390/genes12010030.
- Pégorier P., Bertignac M., Chentli I., Nguefack Ngoune V., Folch G., Jabado-Michaloud J., Hadi-Saljoqi S., Giudicelli V., Duroux P., Lefranc M.-P. and Kossida S. IMGT® biocuration and comparative study of the T cell Receptor beta locus of veterinary species based on Homo sapiens TRB. Front. Immunol., 2020b May 5;11:821doi: 10.3389/fimmu.2020.00821.


