Research teams
Genome dynamics department
Non-coding RNA, epigenetics and genome stability

Understanding the Interactions Between the Genome and Its Transposable Elements
Sequencing of eukaryotic genomes has revealed that genes represent only a small fraction of the genome. Most of it consists of intergenic regions and repeated sequences—such as satellites, microsatellites, and transposable elements (TEs). In humans, for instance, nearly 45% of the genome is derived from these TEs.
These DNA sequences, when functional, can move and multiply within the genome. As true genetic parasites, they exploit the cellular machinery of their host to ensure their propagation, while simultaneously threatening the integrity of its genome. Throughout evolution, organisms have developed and selected a diversity of defense mechanisms to contain TE mobility at a level compatible with their survival.
In the germline, where genetic stability is essential, RNA interference mechanisms act to repress TEs through two main modes of action:
• By blocking the transcription of TEs through epigenetic marks such as DNA methylation or histone modifications: this is referred to as TGS (Transcriptional Gene Silencing).
• By inducing the degradation of TE transcripts: this is referred to as PTGS (Post-Transcriptional Gene Silencing).
A constant dialogue exists between TEs and the host genome, maintaining a dynamic equilibrium. Thus, modern genomes reflect a long evolutionary history shaped by interactions, selections, and genetic drift that have molded their architecture. However, this equilibrium is fragile: the genome remains a dynamic system constantly subjected to constraints. Its stability results from an unstable evolutionary compromise, the disruption of which can lead to pathologies or sterility, particularly when the germline is affected.
Our Research
Our team studies the molecular mechanisms that illustrate the coevolution between transposable elements and their hosts. We aim to understand:
• how TE activity is regulated;
• how TEs multiply while preserving genome integrity;
• how their activities can modify genome structure and regulation;
• how these interactions influence host adaptation to various environmental stresses (heat, oxidative stress, etc.).
Toward an Integrated View of the Genomic Host–Parasite Conflict
Our team’s project is part of an interdisciplinary approach combining genetics, comparative genomics, biochemistry, bioinformatics, advanced imaging, and, more recently, applied mathematics. This diversity of expertise is a true strength in addressing complex and innovative questions at the interface of several disciplines. Our research program thus relies on an integrative strategy that combines precise biological experimentation with large-scale data modeling and analysis.
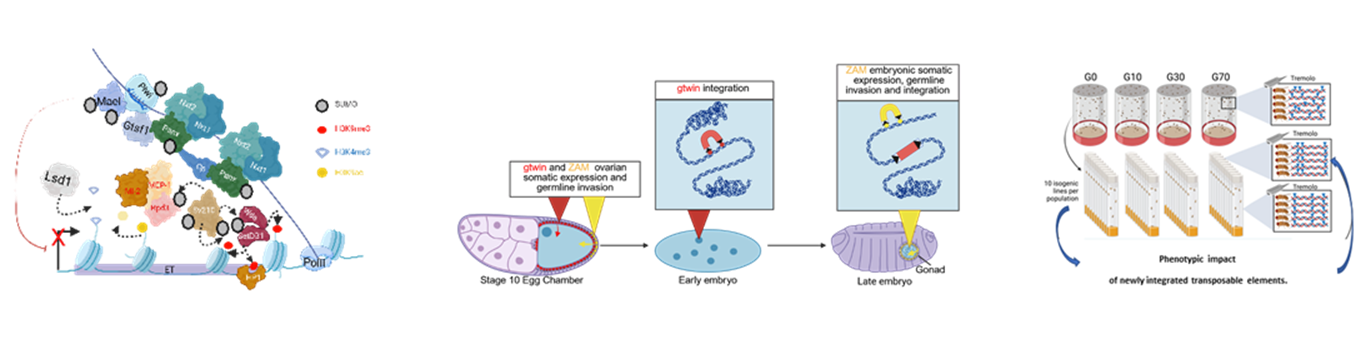
Piwi-interacting RNAs (piRNAs), small non-coding RNAs, once associated with the nuclear Piwi protein (a member of the Argonaute subfamily), recognize and bind to nascent transcripts of transposable elements, thereby triggering a transcriptional repression process. In the female gonad of Drosophila, numerous cofactors involved in this pathway have been identified. These can be grouped into two major functional classes (Figure 1).
The first class includes the core components of the Piwi/piRNA complex, notably Gtsf1/Arx and Maelstrom, as well as the proteins of the SFiNX complex (Panx, Nxf2, Nxt1, Ctp) (Figure 1). The second class includes factors capable of remodeling chromatin to inhibit the transcriptional activity of loci containing transposable elements. Among these are enzymes referred to as “writers,” “erasers,” or “remodelers.” For example, LSD1 (a histone demethylase) and Rpd3 (a histone deacetylase) remove activating histone marks (H3K4me2 and H3K9ac, respectively). SetDB1/Egg, a key writer, catalyzes the trimethylation of H3K9 (H3K9me3), leading to the recruitment of HP1, a protein that stabilizes repressive chromatin.
Recently, we highlighted the involvement of the dMec chromatin remodeling complex, including the ATPase Mi-2 and MEP-1, in the control of transposable elements (Mugat et al., 2020; https://doi.org/10.1038/s41467-020-16635-5).
Despite the identification of many cofactors involved in piRNA-mediated transcriptional repression, understanding their spatiotemporal mode of action remains incomplete. In particular, it is essential to elucidate how these factors dynamically interact at the transcriptional level over time to initiate and maintain a repressive state.
To address this, we are developing approaches that combine real-time imaging and modeling to directly observe the effect of piRNAs on transcriptional dynamics.
Project 2:Understanding the Selectivity and Temporality of Transposable Element Insertion in Drosophila melanogaster
In Drosophila melanogaster, as in most eukaryotes, the genome contains a great diversity of transposable element families. Even when sequences that remain capable of transposition share similar transposition mechanisms, we have shown that each of them exhibits a specific expression profile during Drosophila development. These elements also differ in their integration dynamics: they insert into different open chromatin domains characterized by distinct epigenetic signatures. These observations therefore reveal strong heterogeneity in insertion behaviors and suggest a fine regulation of interactions between transposable elements and their chromatin environment (Varoqui et al., 2025; https://doi.org/10.1093/nar/gkaf516) (Figure 2).
These idiosyncratic expression and integration profiles appear to result from processes of selection and/or genetic drift that have led each transposable element family to occupy specific genomic niches and adopt their own insertion dynamics.
Our project aims to elucidate the molecular determinants of this selectivity of expression and integration. We seek to understand how elements within the same family recognize distinct genomic targets and which host proteins contribute to these differential patterns of expression and integration. To achieve this, we are developing innovative microscopy and biochemical approaches.
Project 3: Impact of New Insertions on the Genome’s Ability to Respond to Stress
Transposable elements are major drivers of genome evolution. However, their involvement in stress response remains poorly understood. To investigate this question, we will use Drosophila melanogaster lines with varying transposable element loads (Barckmann et al., 2018; doi: 10.1093/nar/gky761; Varoqui et al., 2025).
These lines were generated using an innovative genetic strategy that allows the mobilization of transposable elements under laboratory conditions while preserving individual fertility. This approach is based on a transient depletion of Piwi, a key factor in the piRNA pathway responsible for transcriptional repression of transposable elements in the female gonad. We have established Drosophila lines with an increasing number of transposable element insertions after 10, 30, and 70 generations of Piwi protein depletion induction (G10, G30, and G70 lines) (Figure 3).
From these lines, we aim to assess the phenotypic consequences of transposable element accumulation, particularly regarding the ability to adapt to different stresses. Specifically, we are testing:
• The thermal stress response, by exposing populations to high or fluctuating temperatures;
• The oxidative stress response, induced by agents such as paraquat, to evaluate whether variations in transposable element load modulate physiological tolerance and survival.
These analyses will determine to what extent transposable elements contribute to the evolutionary dynamics of genomes and to the adaptive capacity of organisms facing changing environmental conditions.


















- Akkouche, A., Mugat, B., Barckmann, B., Varela-Chavez, C., Li, B., Raffel, R., Pélisson, A., and Chambeyron, S. (2017). Piwi Is Required during Drosophila Embryogenesis to License Dual-Strand piRNA Clusters for Transposon Repression in Adult Ovaries. Mol. Cell 66, 411-419.e4. https://doi.org/10.1016/j.molcel.2017.03.017.
- Mohamed, M., Sabot, F., Varoqui, M., Mugat, B., Audouin, K., Pélisson, A., Fiston-Lavier, A.-S., and Chambeyron, S. (2023). TrEMOLO: accurate transposable element allele frequency estimation using long-read sequencing data combining assembly and mapping-based approaches. Genome Biol 24, 1–20. https://doi.org/10.1186/s13059-023-02911-2.
- Mohamed, M., Dang, N.T.-M., Ogyama, Y., Burlet, N., Mugat, B., Boulesteix, M., Mérel, V., Veber, P., Salces-Ortiz, J., Severac, D., et al. (2020). A Transposon Story: From TE Content to TE Dynamic Invasion of Drosophila Genomes Using the Single-Molecule Sequencing Technology from Oxford Nanopore. Cells 9, 1776. https://doi.org/10.3390/cells9081776.
- Mugat, B., Nicot, S., Varela-Chavez, C., Jourdan, C., Sato, K., Basyuk, E., Juge, F., Siomi, M.C., Pélisson, A., and Chambeyron, S. (2020). The Mi-2 nucleosome remodeler and the Rpd3 histone deacetylase are involved in piRNA-guided heterochromatin formation. Nat Commun 11, 2818. https://doi.org/10.1038/s41467-020-16635-5.
- Varoqui, M., Mohamed, M., Mugat, B., Gourion, D., Lemoine, M., Pélisson, A., Grimaud, C., and Chambeyron, S. (2025). Temporal and spatial niche partitioning in a retrotransposon community of the Drosophila melanogaster genome. Nucleic Acids Res 53, gkaf516. https://doi.org/10.1093/nar/gkaf516.
- Ethève, N.B., Chambeyron, S., and Bantignies, F. (2024). Chapitre 2 – Les acteurs moléculaires de l’information épigénétique (ISTE Group).
- Vitte, C., Chambeyron, S., and Vieira, C. (2024). Chapitre 3 – Épigénétique et éléments transposables (ISTE Group).







