Research teams
Genome dynamics department
Gene regulation

The Gene Regulation Lab at the Institut de Génétique Humaine investigates the molecular mechanisms that control gene expression, RNA metabolism, and genome stability in human cells. We are particularly interested in how RNA molecules and the protein complexes that process them shape chromatin architecture, modulate transcription, and safeguard genome integrity. Our recent research has focused on three interconnected areas: nuclear RNA surveillance, co-transcriptional regulation of microRNAs, and transcription-associated genome instability. Together, these projects provide a comprehensive view of the dynamic interplay between RNA, chromatin, and gene regulation.
The Gene Regulation Lab seeks to uncover fundamental principles of RNA biology and gene regulation, connecting molecular mechanisms to genome organization and stability. By combining proteomics, genomics, and computational approaches, we aim to map the networks that integrate RNA processing, chromatin structure, and transcriptional control. Our ultimate goal is to understand how these processes are coordinated in healthy cells and how their dysregulation contributes to disease. We are committed to fostering a collaborative and innovative research environment, training the next generation of scientists, and translating mechanistic insights into broader understanding of human genetics and molecular medicine.
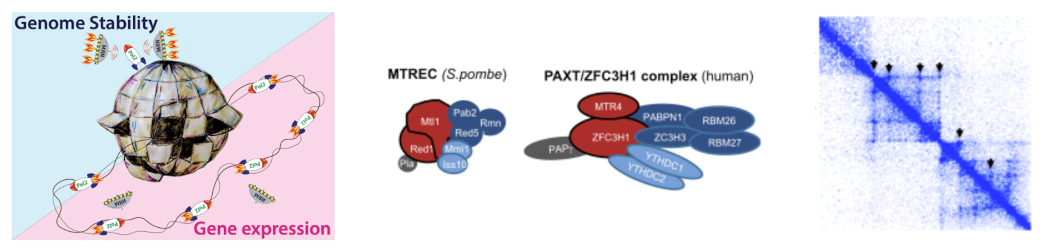
1. Nuclear ncRNA Surveillance: Mechanisms and Impact on Transcription and Genome Organization
Cells transcribe vast portions of the genome, generating not only protein-coding mRNAs but also numerous non-coding RNAs (ncRNAs), including promoter upstream transcripts (PROMPTs) and enhancer RNAs (eRNAs). These unstable RNAs are tightly regulated by nuclear RNA surveillance complexes to prevent spurious gene activation and preserve chromatin organization.
Building upon earlier work implicating nuclear RNA surveillance complexes in the silencing of HIV-1 transcription (Contreras et al., PLoS Pathogens, 2018), we have significantly expanded our understanding of the players and processes that connect RNA turnover with gene regulation and 3D genome structure.
PAXT Complex and ncRNA Degradation. Pervasive transcription produces a wide range of unstable RNAs, including enhancer RNAs (eRNAs) and promoter upstream transcripts (PROMPTs), which are selectively degraded by the nuclear exosome. Recruitment of the exosome to target RNAs is mediated by adaptor complexes such as NEXTand PAXT. Through proteomics, we identified novel components of human PAXT, including homologs of the fission yeast MTREC complex. Notably, we discovered poly(A) polymerase gamma (PAPγ) as a new PAXT subunit.
Using ChIP-seq, we showed that PAXT components (ZFC3H1, RBM27, PAPγ) localize to transcription start sites (TSSs) and active enhancers. Loss of ZFC3H1 impaired PAXT recruitment and resulted in the accumulation of PROMPTs. Similarly, PAPγ depletion led to increased ncRNA levels. We also demonstrated that PAXT facilitates polyadenylation of PROMPTs via PAPγ, ZFC3H1, and MTR4. These findings illuminate how PAXT directly engages with genomic loci to control ncRNA stability
(Contreras et al.,Nature Communications, 2023).
Nuclear Exosome Targeting and 3D Genome Architecture. Recent evidence suggests that ncRNAs contribute to shaping enhancer–promoter contacts, beyond the well-known roles of cohesin and insulator factors. We investigated whether nuclear exosome targeting complexes influence genome folding by integrating ChIP-seq data for PAXT (ZFC3H1), NEXT (ZCCHC8), and MTR4 with Hi-C–based 3D chromatin maps.
We found that exosome-targeting factors localize preferentially to enhancer–promoter loop anchor points. Depleting PAXT, NEXT, or MTR4 led to increased accumulation of ncRNAs and enhanced cohesin binding at these sites. MTR4 depletion specifically altered loop extrusion dynamics, increasing contacts at loop anchors while reducing intraloop interactions. These findings reveal that RNA decay pathways actively shape 3D genome structure by regulating ncRNA abundance at architectural elements (Akkawi et al., in revision; bioRxiv doi: 10.1101/2025.08.25.671287).
Taken together, our findings suggest that the abundance of ncRNAs at specific 3D interaction sites can determine enhancer-promoter contacts and influence gene expression. Our future projects aim to understand how this may be regulated at specific sites, to specifically fine-tune gene expression.
We have further investigated a major partner of MTR4 helicase, NRDE2. NRDE2 has been implicated in the regulation of splicing, particularly retention of weak introns, in humans. However, its impact on gene expression has not been extensively studied. Our proteomic analysis revealed that NRDE2 interacts with elongating RNA Polymerase II and transcription-associated complexes such as P-TEFb, PAF, and Integrator. Its depletion affects gene expression genome-wide, largely independent of splicing, pointing to a broad regulatory role in co-transcriptional RNA metabolism and gene regulation (Srbic et al, IJMS, 2025).
YTHDC1 and Heat-Induced Transcriptional Reprogramming. We also investigated YTHDC1, a chromatin-associated m6A reader identified in PAXT proteomics. In collaboration with André Verdel (ABI, Grenoble) and colleagues, we demonstrated that YTHDC1 regulates transcription and splicing of heat shock genes. During heat stress, YTHDC1 dynamically relocates to stress bodies and binds m6A-modified SATIII RNAs, coordinating gene expression reprogramming (Timcheva et al., Cell Reports, 2022).
2. Regulation of co-transcriptional miRNA processing by RNA binding proteins
MicroRNAs (miRNAs) are essential post-transcriptional regulators of gene expression, and their biogenesis is tightly coordinated with transcription. Our lab has characterized the RNA-binding protein NF90/NF110 as a selective regulator of intronic pri-miRNA processing. NF90 binds specific pri-miRNAs based on RNA structural features, inhibiting their cleavage by the Microprocessor complex.
Beyond miRNA maturation, NF90 interacts with components of the RNA-induced silencing complex (RISC), including Mov10 and Ago2, modulating post-transcriptional gene silencing. These findings reveal NF90 as a multifunctional RNA-binding protein that integrates transcriptional and post-transcriptional regulation to fine-tune gene expression.
Selective Modulation of Pri-miRNAs by NF90. We previously showed that NF90/NF110 regulates Dicer expression through modulation of intronic pri-miRNA processing (Barbier et al., Cell Research, 2018). Expanding this work, we characterized NF90’s broader role in RNA metabolism in collaboration with Shuji Sakamoto (Kochi University, Japan) and William Ritchie (IGH).
We next identified a subset of intronic pri-miRNAs whose processing is inhibited by NF90 via competition with Microprocessor. Binding is determined by stem stability: destabilizing the stem region abrogates NF90 binding. These results underscore how structural RNA features mediate selective processing control (Grasso et al., Nucleic Acids Research, 2020).
NF90 Interacts with RISC Components and Modulates Ago2 Association. Proteomics revealed NF90 association with Mov10 and RISC components. We showed that NF90 affects Ago2–mRNA binding, extending its role to RNA silencing and post-transcriptional control (Grasso et al., BMC Biology, 2022).
NF90 as a Multifunctional Regulator in RNA Biology. In a recent review, we synthesized data supporting NF90’s broad roles in RNA processing, stability, and translation (Grasso & Kiernan, IJMS, 2022).
3. Transcription-Associated Genome Instability
Transcription, while essential, poses risks to genome integrity. We explored this paradox using HIV-1 as a model of highly active transcription. Using the Proteomics of Isolated Chromatin Fragments (PICH) approach, we identified MDC1 and the MRN complex (MRE11, RAD50, NBS1) as key factors recruited to transcribed loci.
ChIP-seq revealed that MRN components localize to active genes in an RNAPII-dependent manner. Depletion of MRN increased RNAPII occupancy and, over time, led to accumulation of single-nucleotide variants across transcribed regions. These results support a model in which MRN dynamically scans active genes to safeguard against transcription-induced DNA damage (Salifou et al., Science Advances, 2021). This study was carried out in collaboration with Jérôme Dejardin (IGH) and Olivier Cuvier (CBI, Toulouse).
This work underscores the delicate balance between transcription and genome stability and highlights how transcription-associated factors protect the genome from instability that could lead to disease.














- C Akkawi, A Heurteau, X Contreras, MHelsmoortel, J. Bossuyt, O Fosseprez, D Dépierre, O Cuvier* and R Kiernan*. Nuclear exosome targeting complexes modulate 3D chromatin Interactions at enhancers/promoters genome-wide. In revision. Preprint https://doi.org/10.1101/2025.08.25.671287
- M Srbic, C Belhaouari, R Raffel, L Lemaire, J Barbier, J Bossuyt, C Akkawi, X Contreras and R Kiernan. NRDE2 Interacts with an Early Transcription Elongation Complex and Widely Impacts Gene Expression. IJMS. 26:9792 doi: 10.3390/ijms26199792.
- X Contreras*, D Depierre*, C Akkawi*, M. Srbic, M Helsmoortel, K Salifou, M LeHars, O Cuvier and R Kiernan*. 2023. PolyA polymerase gamma associates with PAXT nuclear exosome to control the stability of PROMPT ncRNAs. Nat Comms 14:6745. doi: 10.1038/s41467-023-42620-9.
- K Timcheva, S Dufour, L Touat-Todeschini, C Burnard, M-C Carpentier, F Chuffart, R Merret, M Helsmoortel, S Ferré, A Grézy, Y Couté, S Rousseaux, S Kochbin, C Vourc’h, C Bousquet-Antonelli, R Kiernan, D Seigneurin-Berny* and A Verdel*. 2022. Chromatin-associated YTHDC1 coordinates heat-induced reprogramming of gene expression. Cell Reports https://doi.org/10.1016/j.celrep.2022.111784
- G Grasso and R Kiernan. 2022. The Polyvalent Role of NF90 in RNA Biology. Int J Mol Sci. 2022 23:13584. doi: 10.3390/ijms23211358
- G Grasso, C Akkawi, C Franckhauser, R Nait-Saidi, M Bello, J Barbier and R Kiernan*. 2022. NF90 Interacts with Components of RISC and Modulates Association of Ago2 with mRNA. BMC Biology 20:194. doi: 10.1186/s12915-022-01384-2.
- G Sanchez, J Barbier, C Elie, R Kiernan and S Rouquier*. 2022. PAF1 facilitates RNA polymerase II ubiquitination by the Elongin A complex through phosphorylation by CDK12. bioRXiv (https://www.biorxiv.org/content/10.1101/2020.09.17.297960v1)
- K Salifou, C Burnard, P Basavarajaiah, G Grasso, M Helsmoortel, V Mac, D Dépierre, C Franckhauser, E Beyne, X Contreras, J Dejardin, S Rouquier, O Cuvier andR Kiernan*. Chromatin-associated MRN complex protects highly transcribing genes from genomic instability. Science Advances 7:Eabb2947. DOI: 10.1126/sciadv.abb2947
- T Higuchi, R Kiernan and S Sakamoto. Characteristic analysis of a secondary structure of primary microRNA bound to a double-strand RNA-binding protein, NF90. Electrophoresis Letters65:13-16.
- G Grasso, T Higuchi, V Mac, J Barbier, M Helsmoortel, C Lorenzo, G Sanchez, M Bello, W Ritchie, S Sakamoto and R Kiernan*. NF90 Modulates Processing of a Subset of Human Pri-miRNAs. Nucl Acids Res. 48: 6874-6888. https://doi.org/10.1093/nar/gkaa386
- K Salifou, R Kiernan and X Contreras. 2019. Nuclear RNA Surveillance complexes control HIV-1 transcription. (French) Med Sci (Paris). 35:113-115. doi: 10.1051/medsci/2019018.
- J Barbier, X Chen, G Sanchez, M Cai, M Helsmoortel, T Higuchi, P Giraud, X Contreras, G Yuan, Z Feng, R Nait-Saidi, O Deas, L Bluy, JG Judde, S Rouquier, W Ritchie, S Sakamoto, D Xie* and R Kiernan*. An NF90/NF110-mediated feedback amplification loop regulates dicer expression and controls ovarian carcinoma progression. Cell Res. 28: 556-571.
- X Contreras, K Salifou, G Sanchez, M Helsmoortel, E Beyne, L Bluy, S Pelletier, E Rousset, S Rouquier and R Kiernan*. 2018. Nuclear RNA surveillance complexes silence HIV-1 transcription. PLoS Pathog. 14(3):e1006950. DOI: 10.1371/journal.ppat.1006950.






