Research teams
Molecular bases of human diseases
Maintenance of genome integrity during DNA replication

Genome integrity is most vulnerable during the S phase of the cell cycle, when cells must accurately duplicate their genetic material before mitosis. This is carried out by thousands of replication forks that progress rapidly along the chromosomes until they converge. Throughout this process, replication forks frequently encounter obstacles that can cause them to stall, a phenomenon known as replication stress. Such stress activates the ATR-CHK1 signaling pathway, which halts cell cycle progression, coordinates fork repair, and prevents chromosome breakage. When this response fails, genomic instability arises, fueling tumorigenesis. Our research investigates how yeast and human cells respond to replication stress, focusing on the mechanisms of transcription-replication conflict resolution, replication fork repair, checkpoint signaling, and the links between replication stress and antitumoral immunity.


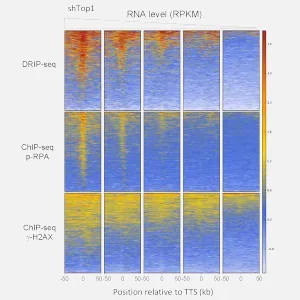
Research in our team is organized into independent yet interconnected projects, each building on our recent advances. These projects use innovative imaging, genomic, and biophysical approaches and benefit from the invaluable technical support of Antoine Barthe, Sandrine Gimenez, Nail Bouzalmad, and Salomé Grassin.
Impact of RNA:DNA hybrids on DNA replication
Lavinia Grasso
A major unresolved question is how R-loops induce replication stress. Our genome-wide studies show that these structures act as polar replication pause sites at the ends of highly expressed genes but do not cause chromosome breaks (Promonet, 2020). We have also identified a distinct class of RNA:DNA hybrids that interfere with the processing and recovery of stalled replication forks (Heuzé, 2023). Using NGS-based and DNA spreading assays, we characterize these “post-replicative” RNA:DNA hybrids to understand their formation and impact. Our working hypothesis is that these hybrids arise as a consequence of replication stress and obstruct fork repair mechanisms.
Replication checkpoint and gene gating
Jonathan Heuzé, Giovanni Nigro, Jérôme Poli
We also study how transcription interferes with replication independently of RNA:DNA hybrids. A likely mechanism involves the accumulation of positive DNA supercoiling when replication and transcription machineries converge, particularly when active genes are tethered to nuclear pores, as in budding yeast. This topological stress may pause replication forks, activate the replication checkpoint, and trigger checkpoint-dependent displacement of RNA polymerase from chromatin, promoting gene release from the nuclear periphery. To test this model, we combine DNA combing, advanced microscopy, and molecular genetics.
Resolution of Top1 cleavage complexes
Benjamin Pardo
Topoisomerase I (Top1) relieves positive DNA supercoiling generated during replication, transcription, and repair by forming transient covalent complexes with DNA, termed Top1 cleavage complexes (Top1ccs). When trapped on DNA, Top1ccs become a major source of damage, but the underlying mechanisms remain poorly defined. Using genome-wide mapping of Top1cc hotspots, DNA breaks, replication forks, and transcription in replicating yeast cells, we investigate how Top1ccs affect genome stability. We also target Top1ccs at defined genomic loci to study their impact on transcription and replication through genomics and single-cell live imaging. This work provides insights relevant to cancer therapies employing Top1 inhibitors.
Chromatin loops, SMCs and topological stress
Solen Faidherbe, Armelle Lengronne
The three-dimensional organization of the genome is a key yet incompletely understood determinant of genome function. Structural Maintenance of Chromosomes (SMC) complexes, including cohesion and condensin, partition chromosomes into large topological domains. We explore how this organization shapes gene expression, DNA replication, and repair using genetic, genomic, and imaging approaches. In particular, we study how cohesin, condensin, and topoisomerases modulate chromatin loops and DNA supercoiling to prevent transcription–replication conflicts.
Origin of cytosolic DNA induced by replication stress
Silpa Mary Johnson, Yea-Lih Lin
The ATR–CHK1 pathway mediates a cell-autonomous response to replication stress, but additional mechanisms signal this stress beyond cell boundaries. We have shown that the processing of stalled replication forks releases small DNA fragments that accumulate in the cytosol, activating the cGAS-STING pathway (Coquel, 2018). This activation triggers type I interferon (IFN) and proinflammatory cytokine production, promoting anticancer responses such as senescence (Técher, 2024) and antitumor immunity. To determine the origin of this cytosolic DNA, we use NGS to compare the sequences of purified cytosolic and replicated DNA from S-phase–synchronized cells treated with hydroxyurea. We also examine the roles of translocases, nucleases, and recombinases in cytosolic DNA generation under various conditions, including oncogene-induced replication stress.
Microfluidic analysis of cytosolic DNA species
Bayan Chami
Characterizing cytosolic DNA has been challenging due to its low abundance and heterogeneous nature. We have developed a microfluidic approach (µLAS) that accurately measures the length and absolute concentration of cytosolic DNA species directly from cell lysates, without extraction or precipitation. We also employ assays to determine the molecular identity of cytosolic nucleic acids—whether double-stranded DNA, single-stranded DNA, or RNA:DNA hybrids—produced under different stress conditions. This strategy provides a powerful means to dissect the interplay between replication stress and inflammation in cancer cells.
Exploiting cytosolic DNA to fight cancer
Diane Amintas, Yea-Lih Lin
We previously demonstrated that inhibition of the cytosolic nuclease TREX1 enhances oncogene-induced senescence by stabilizing cytosolic DNA and triggering a type I IFN response in an MRE11-dependent manner (Técher, 2024). We now seek to determine how TREX1 inhibition suppresses tumor growth in vivo and promotes tumor rejection in immunocompetent mice, in collaboration with Laurent Gros (IRCM).
Developing non-viral vectors for gene therapy
Pauline de Oliveira, Nadege Gaborit
As part of the REPLIBIO project, funded by the PEPR BBTI program (France 2030), we are developing novel non-integrative vectors for gene therapy based on human replication origins. This work extends the pioneering Vectori project initiated by Marcel Méchali and is carried out in close collaboration with Genethon and three research teams from IGH and CBS (Montpellier).






































- Coquel F, Ho SZ, Tsai KC, Yang CY, Aze A, Devin J, Chang TH, Kong-Hap M, Bioteau A, Moreaux J, Maiorano D, Pourquier P, Yang WC*, Lin YL* and Pasero P* (2025) Synergistic effect of inhibiting CHK2 and DNA replication on cancer cell growth. eLife, 13, RP104718
- Técher H, Gopaul D, Heuzé J, Bouzalmad N, Leray B, Vernet A, Mettling C, Moreaux J, Pasero P* and Lin YL* (2024) MRE11 and TREX1 control senescence by coordinating replication stress and interferon signaling. Nat Commun, 15, 5423
- Heuzé J#, Kemiha S#, Barthe A, Torán Vilarrubias A, Aouadi E, Aiello U, Libri D, Lin Y-L*, Lengronne A*, Poli J* and Pasero P* (2023) RNase H2 degrades toxic RNA:DNA hybrids behind stalled forks to promote replication restart. EMBO J, 2022113104
- Forey R, Barthe A, Tittel-Elmer M, Wery M, Barrault MB, Ducrot C, Seeber A, Krietenstein N, Szachnowski U, Skrzypczak M, Ginalski K, Rowicka M, Cobb J, Rando OJ, Soutourina J, Dubrana K, Gasser SM, Morillon A, Pasero P, Lengronne A* and Poli J* (2021) A role for the Mre11–Rad50–Xrs2 complex in gene expression and chromosome organization. Mol Cell, 81,183-197
- Promonet A#, Padioleau I#, Liu Y#, Sanz L, Biernacka A, Schmitz AL, Skrzypczak M, Sarrazin A, Mettling C, Rowicka M, Ginalski K, Chedin F, Chen CL*, Lin YL* and Pasero P* (2020) Topoisomerase 1 prevents replication stress at R-loop-enriched transcription termination sites. Nat Commun, 11, 3940
- Pardo B*, Moriel-Carretero M, Vicat T, Aguilera A* and Pasero P (2020) Homologous recombination and Mus81 promote replication completion in response to replication blockage. EMBO Rep, 21, e49367
- Forey R#, Poveda A#, Sharma S, Padioleau I, Renard C, Lambert R, Skrzypczak M, Ginalski K, Lengronne A, Chabes A, Pardo B* and Pasero P* (2020) Mec1 is activated at the onset of normal S phase by low dNTP pools impeding DNA replication. Mol Cell, 78, 396-410 Recommended by F1000
- Delamarre A, Barthe A, de la Roche Saint‐André C, Luciano P, Forey R, Padioleau I, Skrzypczak M, Ginalski K, Géli V*, Pasero P* and Lengronne A* (2020) MRX increases chromatin accessibility at stalled replication forks to promote nascent DNA resection and cohesin loading. Mol Cell, 77, 395-410
- Bianco JN, Bergoglio V, Lin YL, Pillaire MJ, Schmitz AL, Gilhodes J, Lusque A, Mazières J, Lacroix-Triki M, Roumeliotis TI, Choudhary J, Moreaux J, Hoffmann JS, Tourrière H* and Pasero P* (2019) Overexpression of Claspin and Timeless protects cancer cells from replication stress in a checkpoint-independent manner. Nat Commun, 10, 910
- Bacal J#, Moriel-Carretero M#, Pardo B, Barthe A, Sharma S, Chabes A, Lengronne A and Pasero P (2018) Mrc1 and Rad9 cooperate to regulate initiation and elongation of DNA replication in response to DNA damage. EMBO J, 37, e99319 News & Views in EMBO J, Recommended by F1000
- Coquel F#, Silva MJ#, Técher H#, Zadorozhny K, Sushma S, Nieminuszczy J, Mettling C, Dardillac E, Barthe A, Schmitz AL, Promonet A, Cribier A, Sarrazin A, Niedzwiedz W, Lopez B, Costanzo V, Krejci L, Chabes A, Benkirane M, Lin YL* and Pasero P* (2018) SAMHD1 acts at stalled replication forks to prevent induction of interferon. Nature, 557, 57-61. Comments in Nature, Nature Reviews Immunology, Cancer Discovery, F1000
- Fang D#, Lengronne A#, Shi D#, Forey R, Skrzypczak M, Ginalski K, Yan C, Wang X, Cao Q, Pasero P* and Lou H* (2017) Dbf4 recruitment by forkhead transcription factors defines an upstream rate-limiting step in determining origin firing timing. Gene Dev, 31, 2405-2415 Recommended by F1000
- Yoshida K, Bacal J, Desmarais D, Padioleau I, Tsaponina O, Chabes A, Pantesco V, Dubois E, Parrinello H, Skrzypczak M, Ginalski K, Lengronne A*, Pasero P* (2014) HDACs act on ribosomal DNA to control the yeast replication program and the competition between origins for limiting initiation factors. Mol Cell, 54, 691-697
- Tittel-Elmer M#, Lengronne A#, Davidson MB, Bacal J, François P, Hohl M, Petrini J, Pasero P* and Cobb JA* (2012) Cohesin association to replication sites depends on Rad50 and promotes fork restart. Mol Cell, 48, 98-108. (*corresponding authors) Recommended by F1000
- Poli J, Tsaponina O, Crabbe L, Keszthelyi A, Pantesco V, Chabes A, Lengronne A* and Pasero P* (2012). dNTP pools determine fork progression and origin usage under replication stress. EMBO J 31, 883-894 (*corresponding authors)
- Crabbé L, Thomas A, Pantesco V, De Vos J, Pasero P* and Lengronne A* (2010) Genomic analysis of replication profiles identifies RFCCtf18 as a key mediator of the replication stress response. Nat Struct Mol Biol, 17, 1391-1397 (* equal contribution)
- Tuduri S, Tourrière H and Pasero P (2010) Defining replication origin efficiency using DNA fiber assays. Chrom Res, 18, 91-102.
- Tuduri S, Crabbé L, Conti C, Tourrière H, Holtgreve-Grez H, Jauch A, Pantesco V, de Vos J, Theillet C, Thomas A, Pommier Y, Tazi J, Coquelle A* and Pasero P* (2009) Topoisomerase 1 suppresses replication stress and genomic instability by preventing interference between replication and transcription. Nat Cell Biol, 11, 1315-1324 (* equal contribution)
- Alabert C, Bianco J and Pasero P (2009) Differential regulation of homologous recombination at DNA breaks and replication forks by the Mrc1 branch of the S-phase checkpoint. EMBO J, 28, 1131-1141
- Zaidi IW, Rabut G, Poveda A, Scheel H, Malmström J, Ulrich H, Hofmann K, Pasero P *, Peter M * and Luke B (2008) Rtt101p and Mms1p form a Cul4Ddb1-like ubiquitin ligase, which promotes replication through damaged DNA. EMBO Rep, 9, 1034-1040
- Luke B, Versini G, Jaquenoud M, Waris Zaidi I, Kurz T, Pintard L, Pasero P* and Peter M* (2006) The cullin Rtt101p promotes replication fork progression through damaged DNA and natural pause sites. Curr Biol, 16, 786-792 (* corresponding authors). (IF 11.0)
- Tourrière H#, Versini G#, Cordon-Preciado V, Alabert C and Pasero P (2005) Mrc1p and Tof1p promote replication fork progression and recovery of independently of Rad53p. Mol Cell, 19, 699-706. (# equal contribution) Recommended by F1000 (IF 15.0)
- Versini G, Comet I, Wu M, Hoopes L, Schwob E and Pasero P (2003) The yeast Sgs1 helicase is differentially required for genomic and ribosomal DNA replication. EMBO J 22, 1939-1949. (IF 10.5)











