Research teams
Genetics, cell biology and development department
Tubulin code

Microtubules (MTs) are essential components of the eukaryotic cytoskeleton built from the polymerisation of α- and β-tubulin heterodimers. They also play various essential roles in cell division, polarity and morphogenesis. In addition, they also provide structural support for cilia, flagella and centrioles. Therefore, despite their structural similarities, these simple polymers perform a remarkably wide range of functions. The major source of their functional diversity lies in the C-terminal tails of tubulins that are exposed at the MT surface and control the recruitment of specific effector proteins, therefore translating molecular diversity into functional specialisation. The molecular diversity of tubulin tails is explained by the “tubulin code” hypothesis which combines the reversible post-translational modifications (PTMs) on tubulin C-terminal tails and the ability of MTs to polymerise from multiple tubulin isotypes (e.g., eight α- and eight β-tubulin isotypes in mammals). Thus, the combination of these different isotypes and PTMs ultimately regulates MT properties and effector binding. The main goal of the team is to identify the enzymes generating these PTMs and to characterize their physiological functions.
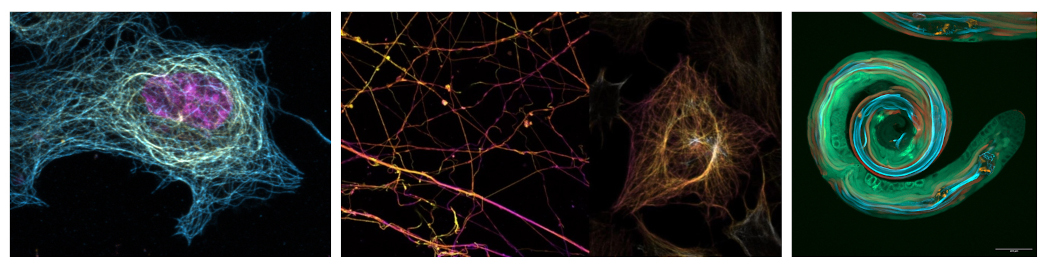
Detyrosination is a widespread tubulin modification consisting in the removal of the C-terminal tyrosine from alpha-tubulin primary chain. This is a reversible modification where the addition of tyrosine back to alpha-tubulin is catalyzed by the Tubulin Tyrosine Ligase TTL. Our lab has recently identified the two classes of enzymes able to cleave the tyrosine from alpha-tubulin, including the Vasohibins VASH1 and VASH2 (Aillaud et al. Science 2017) as well as the metalloprotease TMCP1/MATCAP (Nicot et al. Sci Adv 2023). Despite detyrosination being discovered almost 60 years ago, its physiological roles are still poorly characterized. In order to tackle this problem, we have developed over the past few years, in collaboration with the team of Lubomir Vezenkov (IBMM, Montpellier), highly potent and specific inhibitors of VASHs (Feral et al. J Med Chem, 2025). On the one hand, we used these inhibitors in lung cancer cell lines to assess the role of detyrosination during EMT (Impheng et al, PNAS, 2026). Our data revealed a link between detyrosination and the protein levels of an important epithelial marker involved in cell-cell adhesion called E-cadherin and we are currently investigating the molecular mechanism in place. On the other hand, one of the main characteristics of neurodegenerative disorders is the loss of microtubule stability, that could be due to the accumulation of detyrosination. Our goal is to apply the newly developed inhibitors in the treatment of neurodegenerative diseases.
New α-tubulin processing enzymes acting in Drosophila germlines
Detyrosination is a post-translational modification associated with stable microtubules, which consists in the removal of the last amino acid from the C-terminus of α-tubulin. Up to now, two families of detyrosinases have been identified in mammalian cells. Nevertheless, these families are not conserved in Drosophila where detyrosination takes place specifically in the male germline. Through a genetic screen we recently identified a third class of detyrosinase acting during fly spermatogenesis. The first part of this project aims at characterizing the function of this enzyme and the role of detyrosination in the male germline. Strikingly, we provide strong evidence that a female-specific isotype of α-tubulin, which carries a terminal phenylalanine, is similarly processed in the female germline, by another enzyme. Based on our recent discovery of the new detyrosinase TMCP1 in mammals (Nicot et al. Sci Adv 2023) and of a new detyrosinase in Drosophila, we propose in a second part a strategy to identify this unknown “dephenylalaninase” and analyze its function. This project will therefore lead to the characterization of two new families of α-tubulin carboxy-peptidases and paves the way for the identification of their putative functional homologs in mammals.
Extending the tubulin code through beta-tubulin processing
Microtubules (MTs) are essential cytoskeletal polymers of alpha- and beta-tubulin that organise cell architecture, drive intracellular transport and morphogenesis, and enable cell division. Their versatility stems from the C-terminal tails of alpha- and beta-tubulins, exposed at the MT surface, which recruit specific effector proteins and motors. The use of distinct tubulin isotypes and various post-translational modifications (PTMs) on these tails establishes a “tubulin code” that translates MT composition into defined functions. Among the best-characterised PTMs is the detyrosination of alpha-tubulin, corresponding to the enzymatic removal of its terminal tyrosine. Defects in this process are linked to cardiomyopathies, cancer, and neurodegeneration, underscoring the importance of tubulin tail processing for MT function. While alpha-tubulin processing has been extensively studied, the regulation of beta-tubulin remains largely unexplored. We recently identified the first enzyme, named TMCP2, that cleaves C-terminal residues of beta-tubulin, generating previously undescribed modifications (Nicot et al. Sci Adv 2023). This project aims at defining TMCP2 substrate specificity, identify proteins that selectively bind to modified beta-tubulins, and determine how these new modifications influence MT behaviour and cellular functions. Using proteomics, advanced microscopy, and genome editing, we will carry out mechanistic studies to link beta-tubulin modifications to specific cellular phenotypes. Finally, analyses in TMCP2 knockout mice will establish the physiological relevance of beta-tubulin processing in vivo. By expanding the tubulin code concept to beta-tubulin, this project will uncover a new layer of cytoskeletal regulation and provide fundamental insights into how MT diversity supports cellular specialisation.














- Impheng, H. et al. Peptide-based covalent inhibitor of tubulin detyrosination promotes mesenchymal-to-epithelial transition in lung cancer cells. Proc Natl Acad Sci U S A. 2026 Jan 6;123(1):e2514990123. doi: 10.1073/pnas.2514990123.
- Simon, M. et al. Real-time FRET assay for monitoring detyrosination by TMCP1 and VASH2. Protein Sci. 2025 Dec;34(12):e70374. doi: 10.1002/pro.70374.
- Feral, A. et al. Development of Low-Nanomolar Covalent Epoxide Inhibitors of Tubulin Detyrosinating Enzymes VASH1&2. Med. Chem. (2025) doi:10.1021/acs.jmedchem.5c00980.
- Corrales, R. M. et al. Tubulin detyrosination shapes Leishmania cytoskeletal architecture and virulence. Natl. Acad. Sci. 122, (2025).
- Nicot, S. et al. A family of carboxypeptidases catalyzing α- and β-tubulin tail processing and deglutamylation. Adv. 9, eadi7838 (2023).
- Rogowski, K., Hached, K., Crozet, C. & Laan, S. van der. Tubulin modifying enzymes as target for the treatment of tau-related diseases. Ther. 218, 107681 (2021).
- Laan, S. van der et al. Evolutionary Divergence of Enzymatic Mechanisms for Tubulin Detyrosination. Cell Reports 29, 4159-4171.e6 (2019).
- Laan, S. van der, Dubra, G. & Rogowski, K. Tubulin glutamylation: a skeleton key for neurodegenerative diseases. Neural Regen Res 14, 1899–1900 (2019).
- Bompard, G. et al. CSAP Acts as a Regulator of TTLL-Mediated Microtubule Glutamylation. Cell Reports 25, 2866-2877.e5 (2018).
- Devambez, I. et al. Identification of DmTTLL5 as a Major Tubulin Glutamylase in the Drosophila Nervous System. Sci Rep-uk 7, 16254 (2017).
- Aillaud, C. et al. Vasohibins/SVBP are tubulin carboxypeptidases (TCPs) that regulate neuron differentiation. Science 358, 1448–1453 (2017).
- Rogowski, K. et al. A Family of Protein-Deglutamylating Enzymes Associated with Neurodegeneration. Cell 143, 564–578 (2010).
- Lacroix, B. et al. Tubulin polyglutamylation stimulates spastin-mediated microtubule severing. J Cell Biology 189, 945–954 (2010).
- Rogowski, K. et al. Evolutionary Divergence of Enzymatic Mechanisms for Posttranslational Polyglycylation. Cell 137, 1076–1087 (2009).
- Janke, C., Rogowski, K. & Dijk, J. van. Polyglutamylation: a fine‐regulator of protein function? Embo Rep 9, 636–641 (2008).
- Dijk, J. van et al. A Targeted Multienzyme Mechanism for Selective Microtubule Polyglutamylation. Mol Cell 26, 437–448 (2007).
- Janke, C. et al. Tubulin Polyglutamylase Enzymes Are Members of the TTL Domain Protein Family. Science 308, 1758–1762 (2005).





