Research teams
Genetics, cell biology and development department
Epigenetic chromatin regulation

Small RNA-directed programmed DNA elimination of transposon-derived sequences
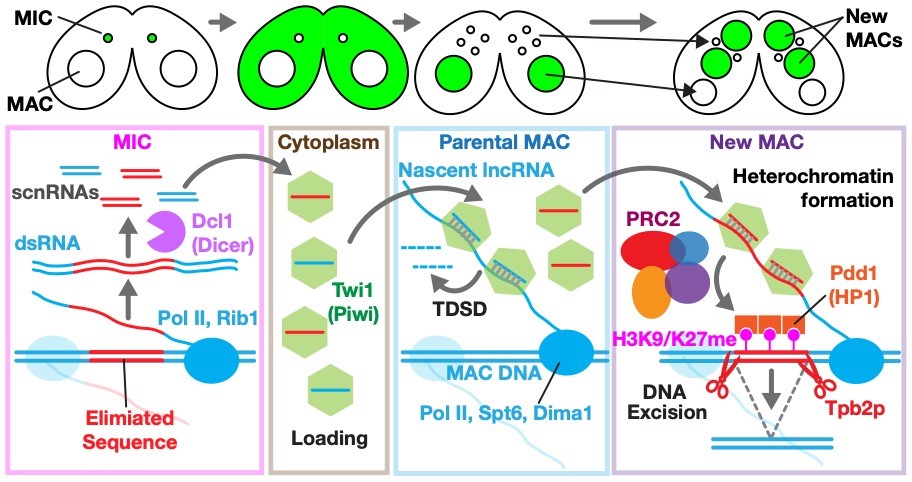
Small RNA-mediated transcriptional gene silencing is a vital defense mechanism shared by diverse eukaryotes from fungi and plants to flies, worms, and mammals. Its primary mission is to keep transposable elements (TEs) in check, preventing these genomic “jumping genes” from causing instability or disease. Small RNAs recognize and silence TEs through precise base-pairing, yet how cells achieve this selectivity without silencing normal genes remains an open question. The ciliated protozoan Tetrahymena offers a remarkable window into this process. During programmed DNA elimination (PDE), it distinguishes TE-derived sequences by comparing its germline and somatic genomes using small RNAs, an elegant example of epigenetic genome regulation and TE–host coevolution. Because DNA elimination can be synchronously triggered on a large scale in the lab, Tetrahymena provides a powerful system for dissecting small RNA-guided chromatin regulation at both genetic and biochemical levels. Using this “tiny, hairy” eukaryote, we aim to uncover how cells: selectively produce small RNAs from TE-related sequences, use them to recognize matching genomic regions, and build the silent chromatin landscapes (heterochromatin) that keep TEs under control.
Small RNA-mediated transcriptional gene silencing paradoxically requires transcription at two distinct steps: first, precursors of small RNAs must be transcribed to initiate the process, and second, the target sites of silencing must themselves be transcribed to interact with the small RNA trigger. The molecular mechanisms underlying these “transcriptions for transcriptional silencing” remain poorly understood. We have reported that transcription of small RNA precursors requires specialized transcription factors and mediator-associated proteins that facilitate initiation from G-rich sequences. Moreover, we found that SUMOylation, a post-translational modification, of the transcription factor Spt6 likely plays an important role in switching the transcriptional state from normal mRNA-producing mode to a specific small RNA–receiving mode. We continue to investigate how these specialized transcriptional machineries function in the production of and interaction with small RNAs, using the identified components as stepping stones for further study.
Regulation of Dual Heterochromatic Modifications in Programmed DNA Elimination
Constitutive heterochromatin, typically found around centromeres and telomeres, is associated with methylation of histone H3 lysine 9 (H3K9me), whereas facultative heterochromatin contains developmentally regulated genes and is marked by H3K27me. Alterations in these modifications can cause diverse diseases: the former maintains genome stability, while the latter preserves cellular identity. H3K9me and H3K27me are generally catalyzed by SUV39 and EZH family methyltransferases, respectively. However, the ciliated protozoan Tetrahymena lacks SUV39, and its EZH enzyme Ezl1, part of the Polycomb Repressive Complex 2 (PRC2), catalyzes both H3K9me and H3K27me during small RNA-directed DNA elimination. We are investigating how PRC2’s dual specificity is regulated, providing insights into the mechanism and evolutionary origins of the division of labor between these heterochromatic pathways.
Cell Sorting-Based Genetic Screening of Programmed DNA Elimination
Investigations into programmed DNA elimination (PDE) in the unicellular ciliate Tetrahymena have revealed how eukaryotes use small noncoding RNAs to regulate transposable elements (TEs) that shape genome organization and integrity. Despite two decades of research, candidate-driven approaches such as gene knockout and protein complex purification have failed to identify many of the molecular factors involved in this process. To further elucidate the small RNA-directed PDE mechanism, this project will employ a high-throughput gene knockdown strategy combined with cell sorting to identify and characterize previously unknown PDE genes through genome-wide forward genetic screening. The study will shed light on the evolution of small RNA-directed TE silencing and advance understanding of conserved principles of small RNA-mediated epigenetic regulation.












- Nagao, K and Mochizuki, K.* (2026) Linking germline telomere removal to global programmed DNA elimination in tetrahymena genome differentiation. eLife. 10.7554/eLife.109351.1.
- Cai, X., Zhai, Z., Noto, T., Dong, G., Wang, X., Liucong, M., Liu, Y., Agreiter, C., Loidl, J., Mochizuki, K., Tian, M.* (2025). A specialized TFIIB is required for transcription of transposon-targeting noncoding RNAs. Nucleic Acids Res. 53. 10.1093/nar/gkaf427.
- Shehzada, S., Noto, T., Saksouk. J., Mochizuki, K.* (2024) A SUMO E3 ligase promotes long non-coding RNA transcription to regulate small RNA-directed DNA elimination. eLife, 13:e95337, doi.org/10.7554/eLife.95337
- Mutazono, M., Noto, T., and Mochizuki, K.* (2019) Diversification of small RNA amplification mechanisms for targeting transposon-related sequences in ciliates. PNAS, 116, 14639-14644. PMID: 31262823
- Noto, T., and Mochizuki, K.* (2018) Small RNA-mediated trans-nuclear and trans-element communications in Tetrahymena DNA Elimination. Curr Biol, 28, 1938-1949. PMID: 29887308
- Kataoka, K., and Mochizuki, K.* (2015) Phosphorylation of an HP1-like protein regulates RNA-bridged heterochromatin body assembly for DNA elimination. Dev Cell 35, 775-788. PMID: 26688337
- Noto, T., Kataoka, K., Suhren, J. H., Hayashi, A., Woolcock, K. J., Gorovsky, M. A. and Mochizuki, K.* (2015) Small RNA-mediated genome-wide trans-recognition network in Tetrahymena DNA elimination. Mol Cell 59, 229-242. PMID: 26095658
- Woehrer, S. L., Aronica, L., Suhren, J. H., Busch, C. J., Noto, T and Mochizuki, K.* (2015) A Tetrahymena Hsp90 co-chaperone promotes siRNA loading by ATP-dependent and ATP-independent mechanisms. The EMBO Journal 34, 559-577. PMID: 25588944
- Schoeberl, U. E., Kurth, H. M., Noto, T. and Mochizuki, K.* (2012) Biased transcription and selective degradation of small RNAs shape the pattern of DNA elimination in Tetrahymena. Genes Dev 26, 1729-1742. PMID: 22855833
- Noto, T., Kurth H. M., Kataoka, K. Aronica, L., Desouza, L.V., Siu, K.W., Pearlman, R.E., Gorovsky, M.A., and Mochizuki, K.* (2010) The Tetrahymena Argonaute-binding protein Giw1p directs a mature Argonaute-siRNA complex to the nucleus. Cell 140, 692-703. PMID: 20211138
- Mochizuki, K., Fine, N. A., Fujisawa, T., and Gorovsky, M. A.* (2002) Analysis of a piwi-related gene implicates small RNAs in genome rearrangement in Tetrahymena. Cell 110, 689-699. PMID: 12297043




