Research teams
Molecular bases of human diseases
Spatial control of genome integrity and oncogenesis (Emerging team)

Over the past decade, our understanding of the mechanisms and functions of cytoplasmic and nuclear membraneless organelles has advanced dramatically, revolutionizing the field of cell biology. Through their spatiotemporal compartmentalization, these mesoscale structures known as biomolecular condensates, contribute to critical cellular processes as cellular metabolism, RNA biogenesis, DNA replication, environmental stress responses, DNA damage control and signal transduction, among others. They are formed through weak, multivalent, cooperative, and site-specific interactions mediated by scaffolding proteins and nucleic acids. In addition, post-translational modifications play a major role in controlling the formation and dissolution of these compartments, thereby conferring reversibility and dynamism to these structures.
The spontaneous formation of nuclear foci in response to deleterious DNA damage is an essential process for locally organizing key cellular factors involved in the detection, signaling, and repair of damaged DNA. These nuclear foci, commonly referred to as biomolecular condensates, concentrate diverse molecules non-stoichiometrically at DNA damage sites. The assembly of these condensates depends on several factors, including the type of DNA lesions, their dynamic composition, and the cell cycle phase during which the damage occurred.
Our team aims to unravel the molecular mechanisms, functions, and regulatory principles governing stress-induced biomolecular condensates, with a particular focus on those involved in the DNA damage response. By integrating cell biology, structural biology, and advanced imaging approaches, we investigate how these condensates assemble, evolve, and function throughout the cell cycle, and how their composition and dynamics influence genome maintenance. A key objective is to understand how alterations in condensate organization contribute to cancer cell survival and proliferation. In collaboration with structural biology partners, we also explore the self-assembly properties of scaffolding proteins and develop innovative strategies to modulate condensate behavior, offering potential new avenues for therapeutic intervention.
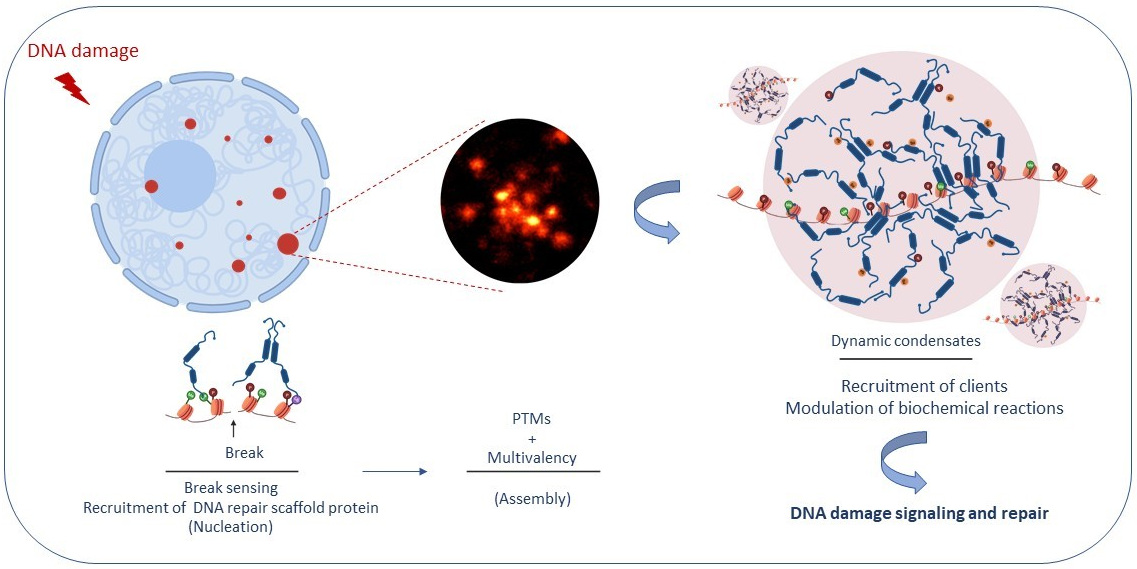
Figure: Role of stress-induced condensates in coordinating specific cellular processes that promote DNA repair and maintain genome integrity.
Our team’s research is organized into two main axes, focusing on the fundamental mechanisms of genome stability and the pivotal role of biomolecular condensates. We investigate how scaffold proteins dynamically assemble into condensates to regulate DNA replication, repair, checkpoint activation, and chromosome segregation. Our projects combine cutting-edge advanced imaging, genomic, and biophysical approaches, supported by strong technical expertise. This work aims to uncover the molecular principles governing these processes and to develop innovative therapeutic strategies, particularly for cancer. Our research thrives on interdisciplinary collaboration among structural biologists, biochemists, and bioinformaticians.
Project 1: TopBP1 condensates: Orchestrating spatial organization of the DNA damage response for genome stability
Elia Moarbess, Sara Mahmoud, Philippe Coulombe
Topoisomerase II-binding protein 1 (TopBP1) is a multivalent scaffold protein that acts as a central hub for DNA replication, repair, transcription, and checkpoint signaling. Our team has shown that, under replication stress, TopBP1 self-assembles into biomolecular condensates, enhancing ATR kinase activity. This phase separation mechanism fine-tunes the DNA damage response and coordinates essential cellular processes. We seek to uncover how the assembly of TopBP1 into biomolecular condensates during S phase modulates nuclear actin dynamics in response to DNA damage, thereby facilitating efficient repair and preserving genomic integrity. Furthermore, we explore how TopBP1 condensates formed during mitosis spatially organize PLK1-dependent phosphorylation events, ensuring the faithful repair of mitotic DNA lesions and safeguarding genome stability.
Project 2: SLX4 condensation and resolution of DNA-protein crosslinks
Philippe Coulombe
SLX4 is a docking platform for structure-specific endonucleases involved in resolving oxidative DNA lesions. We study how SLX4 self-assembly orchestrates the recruitment of repair factors to remove DNA–protein crosslinks generated by oxidative stress. By examining the spatial organization and functional dynamics of SLX4 condensates, we seek to uncover the molecular mechanisms that ensure genome maintenance and cellular stress responses.
Project 3: KIF2C self-assembly and kinetochore regulation during mitosis
Anthony Ibrahim
Accurate chromosome segregation during mitosis is essential for genome stability. This project focuses on KIF2C, a microtubule depolymerase that forms membraneless condensates at kinetochores. We aim to understand how KIF2C assembles into condensates, identify its mitotic binding partners, and determine how these condensates control spindle organization, microtubule-kinetochore attachments, and the spindle assembly checkpoint. Using advanced imaging, structural biology, and optogenetics, we will uncover the mechanisms by which KIF2C ensures mitotic fidelity and provides insights into chromosome segregation defects associated with cancer. This research is conducted in close partnership with Sophie Zinn-Justin’s team (I2BC, Paris)
Project 4: Characterizing the condensation of DNA replication initiator proteins
Lina-Marie Briu
Within the REPLIBIO project, supported by the PEPR BBTI program (France 2030), we are investigating how the condensation of DNA replication initiator proteins enhances the performance of autonomously replicating vectors for gene therapy applications. This work extends the pioneering Vectori project initiated by Marcel Méchali and is carried out in close collaboration with Genethon and three research teams from IGH and CBS (Montpellier).
Project 5: Breaking the condensate code: Targeting biomolecular assemblies for cancer treatment innovation
Elia Moarbess, Philippe Coulombe
Biomolecular condensates play a crucial role in tumor development and progression. These dynamic and reversible assemblies, driven by intrinsically disordered regions (IDRs) in proteins, are emerging as key regulators of cellular stress responses—including the DNA damage response. Among these, TopBP1 condensates are essential for cancer cells, as they amplify ATR/Chk1 signaling to mitigate replication stress and maintain genome stability. We are developing strategies to disrupt TopBP1 condensate formation by targeting its disordered ATR activation domain (AAD) using nanobodies and RF diffusion-based artificial binders, in collaboration with Joseph M. Rogers (University of Copenhagen). By inhibiting condensate assembly, we aim to impair ATR signaling, sensitize cancer cells to chemotherapy, and overcome treatment resistance, particularly in colorectal cancer. This project is carried out in partnership with Sophie Zinn-Justin (I2BC, Paris) and Céline Gogora (IRCM).
















- Morano L.; Vezzio-Vié N., Aissanou A; Hodroj D.; Garambois V. Fauvre A., Promonet A., Egger T., Bordignon B., hassen-khodja C., Fiachetti S., Basbous J*., Gongora C*., Constantinou A. (2025) Shining Light on Drug Discovery: Optogenetic Screening for TopBP1 Biomolecular Condensate Inhibitors. NAR Cancer. 2025 Nov 3;7(4) *Co-corresponding authors
- Morano L., Vezzio-Vié N., Aissanou A., Egger T., Aze A., Fiachetti S., Seitz H., Milazzo L.A., Garambois V., Bonnefoy N., Gongora C*., Constantinou A., Basbous J*. (2025) TopBP1 biomolecular condensates: a new therapeutic target in advanced-stage colorectal cancer. 2025 Oct 21;14:RP106196. * Co-corresponding authors.
- Skobelkina A., Julien M., Jeannin S., Miron S., Egger T., Chaaban R., Alghoul E., Bouvignies G., Ghouil R., Friel C., Busso D., Theillet FX., Le Bars R., Carreira A., Constantinou A., Basbous J*., Zinn-Justin S*. (2025) KIF2C-induced nuclear condensation concentrates PLK1 and phosphorylated BRCA2 at the kinetochore microtubules in mitosis. Nucleic Acids Res. 2025 Jun 6;53(11):gkaf476. * Co-corresponding authors.
- Egger T., Morano L., Blanchard MP., Basbous J*., Constantinou A. (2024) Spatial organization and functions of Chk1 activation by TopBP1 biomolecular condensates. Cell Rep. 2024 Apr 23;43(4):114064. * Corresponding authors.
- AlGhoul E., Basbous J., Constantinou A. (2023) compartmentalization of DNA damage response: Mechanisms and functions. DNA Repair (Amst). 2023 Aug;128:103524.
- AlGhoul E., Paloni M., Takedachi A., Urbach S., Barducci A., Gaillard PH., Basbous J*., Constantinou A*. (2023) Compartmentalization of the SUMO/RNF4 pathway by SLX4 to drive DNA repair. Mol Cell. 2023 May 18;83(10):1640-1658.e9. doi: 10.1016/j.molcel.2023.03.021. * Co-corresponding authors.
- AlGhoul E., Basbous J.*, Constantinou A*. (2021) An optogenetic proximity labeling approach to probe the composition of inducible biomolecular condensates in cultured cells. STAR Protoc. Jul 22;2(3):100677.* Co-corresponding authors.
- Frattini C., Promonet A., AlGhoul E., Vidal-Eychenié S., Lamarque V., Blanchard MP., Urbach S., Basbous J.* and Constantinou A*. (2021) TopBP1 assembles nuclear condensates to switch on ATR signaling. Mol Cell. 2021 Mar 18;81(6):1231-1245.e8. * Co-corresponding authors.
- Basbous J.*, Aze A., Chaloin L., Lebdy R., Hodroj D., Ribeyre C., Larroque M., Shepard C., Kim B., Pruvost A., Moreaux J., Maiorano D., Mechali M. Constantinou A*. (2020) Dihydropyrimidinase protects from DNA replication stress caused by cytotoxic metabolites. Nucleic Acids Res. 2020 Feb 28;48(4):1886-1904. * Co-corresponding authors.
- Basbous J. and Constantinou A. (2019) A tumor suppressive DNA translocase named FANCM. Crit Rev Biochem Mol Biol. 2019 Feb;54(1):27-40.





