Research teams
Genome dynamics department
Biology of repetitive sequences

Our lab studies how repetitive regions of the mammalian genome are organized, maintained, and used by the cell. We focus especially on heterochromatin forming DNA repeats, including chromosomal domains like telomeres and pericentromeres, that are essential for genome stability but still not fully understood at the mechanistic level. A central theme of our work is that these regions are not simply inert or repressive structures. Instead, they are dynamic chromatin environments that can influence replication, repair, transcription, nuclear organization, and chromosome inheritance.
One current line of research examines how heterochromatin controls genome function at telomeres. Our work has shown that the chromatin state of telomeres can actively shape how chromosome ends are maintained, including in contexts linked to replication stress and recombination-based telomere elongation. More broadly, we aim to understand how heterochromatin is assembled, how its function is linked with nuclear architecture, and how its protein effectors regulate chromosome function in development and disease.
A second major focus of the lab is telomere architecture and end protection. We recently identified TELS1, a factor that stabilizes telomere loops in embryonic stem cells, and this opened the way to directly test long-standing models of telomere protection in mammals. We are now exploring how telomere loops are formed and regulated, how chromosome ends are protected during early development, and how cells can sometimes tolerate linear telomeres without triggering aberrant repair. Through these studies, we aim to uncover fundamental principles of telomere biology and genome maintenance.
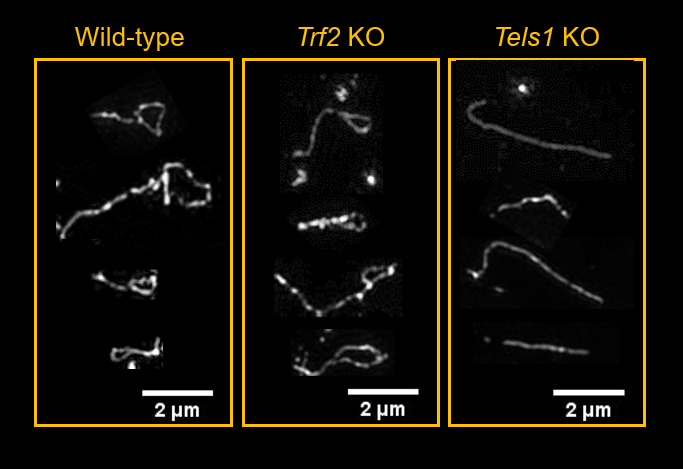
DDR signaling at chromosomes-ends during pre-implantation development
Yen Huyhn, Chiara Gino, Raoudha Fadhloun, Karim Mesbah, Mathieu Tardat
A major ongoing project in the lab is to understand the role of TELS1 and telomere loop dynamics during early development. We are investigating how embryonic stem cells can tolerate largely linear telomeres, why this differs from differentiated cells, and which molecular pathways prevent these chromosome ends from being aberrantly repaired. We are also analyzing a Tels1 knockout mouse model to determine the physiological role of TELS1 in vivo, and studying telomere maintenance at the transition between pluripotent and totipotent-like states, including mechanisms of telomere elongation in very early embryonic contexts.
Biophysical analysis of t-loop formation by TELS1
Mathieu Tardat
In parallel, we are carrying out mechanistic studies of TELS1-dependent telomere loop formation. Using purified components and biophysical approaches in collaboration with the Robert’s lab, we aim to understand how TELS1 promotes t-loop formation. These experiments are designed to move from cellular observations to a molecular description of how telomere architecture is controlled.
Understanding the molecular basis of LAD tethering
Lise Dauban
Another developing research axis focuses on heterochromatin organization at the nuclear periphery. We are studying how large chromosomal regions called lamina-associated domains are targeted to the nuclear lamina, which DNA elements are sufficient to confer this behavior, and which proteins mediate the connection between chromatin and the nuclear envelope. This work combines genomics, chromatin proteomics, and genetic screening approaches to identify the principles governing large-scale heterochromatin positioning in the nucleus.
Heterochromatin flavors
Sandrine Sauzet
Finally, we are investigating how heterochromatin forms at distinct DNA repeats. Our previous work identified several ubiquitin ligases within heterochromatin, and we are now developing the tools needed to define their molecular role more precisely. This project is part of a broader effort to understand how chromatin-associated enzymes and effectors shape distinct heterochromatin states and influence genome stability.
















- TELS1 stabilizes t-loops independently of TRF2 and controls telomere length in pluripotent cells. Joey Dufourd, Thi Hai Yen Huynh, Sandrine Sauzet, Quynh Huong Lai, Francesco Abagnale, Sihem Zitouni, Marie-Josèphe Giraud-Panis, Serge Urbach, Eric Gilson, Mathieu Tardat, Jérôme Déjardin. Cell Rep 2025. Sep 23;44(9):116260. doi: 10.1016/j.celrep.2025.116260. PMID: 40934081 DOI: 10.1016/j.celrep.2025.116260
- SETDB1NSD-dependent H3K9me3/H3K36me3 dual heterochromatin maintains gene expression profiles by bookmarking poised enhancers. Barral A, Pozo G, Ducrot L, Papadopoulos GL, Sauzet S, Oldfield AJ, Cavalli G, Déjardin J. Mol Cell 2022 Jan 17:S1097-2765(21)01142-4.
- SETDB1-dependent heterochromatin stimulates alternative lengthening of telomeres. Gauchier M, Kan S, Barral A, Sauzet S, Agirre E, Bonnell E, Saksouk N, Barth TK, Ide S, Urbach S, Wellinger RJ, Luco RF, Imhof A, Déjardin J. Science Adv 2019 May 8;5(5):eaav3673. doi: 10.1126/sciadv.aav3673. eCollection 2019 May. PMID: 31086817
- Nuclear-receptor-mediated telomere insertion leads to genome instability in ALT cancers. Marzec P, Armenise C, Pérot G, Roumelioti FM, Basyuk E, Gagos S, Chibon F, Déjardin J. Cell 2015 Feb 26;160(5):913-927. doi: 10.1016/j.cell.2015.01.044. PMID: 25723166
- Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Saksouk N, Barth TK, Ziegler-Birling C, Olova N, Nowak A, Rey E, Mateos-Langerak J, Urbach S, Reik W, Torres-Padilla ME, Imhof A, Déjardin J, Simboeck E. Mol Cell 2014 Nov 20;56(4):580-94. doi: 10.1016/j.molcel.2014.10.001. PMID: 25457167





