Research teams
Genetics, cell biology and development department
Systemic impact of small regulatory RNAs

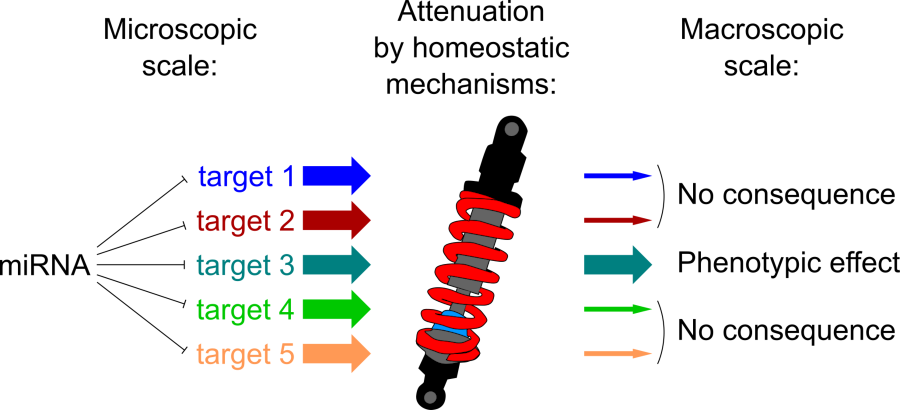
MicroRNAs (“miRNAs”) are small post-transcriptional regulators. The function of these small RNAs in animals has been well characterized at a molecular level, but their role is less well known at the macroscopic scale: how could miRNAs have any biological function if they repress most of their targets less than 2-fold (while inter-individual gene expression fluctuation typically exceeds 2-fold, and is buffered by homeostatic mechanisms)?
According to the current dogma, each miRNA regulates tens or hundreds of targets, yet several observations suggest miRNAs have a much weaker impact on animal biology. Our recent work also suggests that both experimental and computational miRNA target identification methods are heavily contaminated with false positives: these false positives may be truly repressed by miRNAs at the molecular scale, but such a small repressive effect fails to translate into a macroscopic phenotype for most genes.
Our work thus suggests that the biological role of miRNAs has been largely over-estimated. We are currently exploring practical consequences of this new theoretical framework, measuring the contribution of individual miRNA/target interactions to global in vivo phenotypes.
More generally, we are proposing a new vision of gene regulation: a regulatory target is not simply a gene that is affected by a regulatory pathway; it is a gene that is affected enough by the pathway – the extent of a measured regulation needs to be confronted to the robustness of biological systems to fluctuations.
Our projects deal with the identification of biologically important miRNA/target interactions. We put a special emphasis on the distinction between interactions defined by molecular biology (e.g., CLIP, CLASH, differential transcriptomics, …) or computational biology (e.g., using comparative genomics), and genetically-validated interactions controlling actual phenotypes at the organism scale. Our lab has a long-standing expertise both in experimental biology and bioinformatics.
Identification of functional targets for the bantam miRNA
Isabelle Busseau, Pauline De Sousa, Élisabeth Houbron
In Drosophila, the bantam miRNA controls obvious phenotypes (growth and viability). Over the years, several targets for that miRNA have been proposed to participate to these phenotypes (genes hid, enabled, tribbles, …) yet a rigorous genetic assessment is still missing. We use CRISPR/Cas9 genome editing to mutate the seed sequence of the bantam miRNA, as well as its predicted binding sites in selected targets, in a complete in vivo setting. The goal is to abolish and restore molecular interactions on demand in order to interrogate their actual phenotypic importance.
Identification of novel TDMD inducers
Élisabeth Houbron
While miRNAs repress long RNAs exhibiting sufficient sequence complementarity, in some cases the interaction triggers the opposite response (with the miRNA being actively degraded while the long RNA is not). That process is called “TDMD” (target-directed microRNA degradation), and it seems to be primarily determined by the geometry of base-pairing between the two RNA molecules. We have predicted computationally some novel TDMD inducer/miRNA pairs, that we are now assessing experimentally for their ability to trigger miRNA degradation.
Correcting miRNA target prediction using the evolutionary history of miRNAs
Flavia Pavan
Comparative genomics is commonly used to predict biologically important miRNA/target interactions (under the assumption that miRNA binding sites must be functionally important if they have been conserved in evolution). But some sequences may be conserved for other, miRNA-independent reasons, while being fortuitously complementary to miRNA seeds. We are therefore exploring the conservation of predicted miRNA binding sites in vertebrate species which have lost a miRNA family, with the goal of identifying miRNA-dependently and miRNA-independently conserved sites.
Exploring the propagation or regulatory signals along regulatory networks
Estelle Rebecq
Regulatory targets for miRNAs, but also for other gene regulators (transcription factors, RNA-binding proteins) are regulated by these molecules. But the regulatory effect may not propagate efficiently to their downstream effectors: network architecture (e.g., negative feedback loops, which can attenuate it; positive feedback loops, which can amplify it) is expected to modulate the regulatory signal. In this project, we aim to quantify the raw effect of regulators, as well as the modulatory effect by the downstream gene network.














- Seitz H (2024) A new perspective on microRNA-guided gene regulation specificity, and its potential generalization to transcription factors and RNA-binding proteins. Nucleic Acids Res 52(16):9360-9368.
- Janati-Idrissi S, de Abreu MR, Guyomar C, de Mello F, Nguyen T, Mechkouri N, Gay S, Montfort J, Gonzalez AA, Abbasi M, Bugeon J, Thermes V, Seitz H, Bobe J (2024) Looking for a needle in a haystack: de novo phenotypic target identification reveals Hippo pathway-mediated miR-202 regulation of egg production. Nucleic Acids Res 52(2):738-754
- Busseau I, Mockly S, Houbron É, Somaï H, Seitz H (2024) Evaluation of microRNA variant maturation prior to genome edition. Biochimie 217:86-94
- Houbron É, Mockly S, Rafasse S, Gros N, Muriaux D, Seitz H (2023) Biochemistry-informed design selects potent siRNAs against SARS-CoV-2. RNA Biol 20(1):272-280.
- Mockly S, Houbron É, Seitz H (2022) A rationalized definition of general tumor suppressor microRNAs excludes miR-34a. Nucleic Acids Res 50(8):4703-4712


Scripts and datasets for each article:
Seitz (2024) Nucleic Acids Research 52(16):9360–9368 :
Scripts and datasets for the preparation of figures.
Busseau et al. (2023) Biochimie S0300-9084(23)00145-1:
Scripts and datasets for the preparation of figures and supplementary figures – 56 KB
Houbron et al. (2023) RNA Biology 20(1):272–280:
Scripts and datasets for the preparation of figures and supplementary figures – 7,2 GB
Mockly et al. (2022) Nucleic Acids Research 50(8):4703–4712 :
Scripts and datasets for the preparation of figures, supplementary figures and supplementary tables – 636Mo
Brechin et al. (2021) RNA 27(2):151-162 :
Hebras et al. (2020) eLife 9:e60862 :
Richard et al. (2020) Autophagy 17(8):1889–1906 :
Pinzón et al. (2019) PLoS Genetics 15(2):e1007915 :
Mockly et Seitz (2019) Methods in Molecular Biology 1970:291–314 :
Scripts and datasets for the preparation of Table 1
Rodríguez-Martínez et al. (2017) Nature Structural and Molecular Biology 24(3):290–299 :
Pinzón et al. (2017) Genome Research 27(2):234–245 :
Scripts and datasets for the preparation of all 6 figures :
Royo et al. (2015) PLoS Genetics 11(10):e1005461 :
Scripts and data for the generation of figures 2D, 3B and 4A



