Research teams
Genetics, cell biology and development department
Cell biology of RNA

Our group has strong interests in gene expression mechanisms, from transcription to translation. While we are interested in the regulation of these processes and their functional consequences, the big question that moves us is to understand how they occur in the context of a living cell.
Indeed, cells are not only the individual units where gene expression takes place, but they are also incredible objects, highly complex and dynamic. If we consider RNA and proteins, a typical cell contains several hundreds of thousands of different molecular species, with some present in millions of copies per cell while others in only few. In order to function with such a high molecular complexity within a crowded environment, cells have developed a remarkable degree of spatial organization, which also allows a high plasticity and dynamics of molecules. It is to get insights into this very fundamental question that we first developed tools to image single mRNAs in live cells (Mol Cell 1998; Curr Biol 2003). With these tools in hands, and others that we developed later (Nat Methods 2006; Nat Methods 2013; Nat Comm 2016; J Cell Biol 2016; Nat Comm 2018; Nat Comm 2021; Nat Protoc 2023), we aim at imaging the basic mechanisms of gene expression directly in living cells, thereby providing a renewed vision of these fundamental processes.
Our strategy is to invest in methodological developments to access and image new facets of gene expression, usually at the levels of single molecules. These developments are mostly focused on imaging RNA metabolism and they are guided by our current scientific questions. We are also keen to contextualize our findings in the context of human health and in particular cancer.
Our group works in three areas: (1) transcriptional regulation; (2) RNA localization and localized translation; (3) assembly of macromolecular complexes with a focus on co-translational events.
1- Imaging transcription in live cells to reveal the mechanisms of gene regulation
It was discovered ~20 years ago that transcription is an inherently stochastic process (Chubb et al., 2006; see our video below), raising a considerable interest in the gene expression field (Pichon, Mol Cell 2018). The fluctuations of promoter activity, also called gene bursting, are attributed to spontaneous and stochastic switches between various promoter states, such as nucleosome occupied or free, bound ot not by TBP, etc. Characterizing these promoter states at the kinetic and molecular levels is of great interest because this gives a direct access to the mechanisms underlying promoter function and regulation. In addition, these spontaneous promoter fluctuations also create cell-to-cell heterogeneities in clonal populations, which can have important functional consequences in some biological systems like HIV-1, where they control viral latency.
We recently developed several optimized RNA imaging tools that are able to track the activity of promoters for extended periods of time and with a sensitivity of a single polymerase (Nat Comm 2016, RNA 2024). This enabled us to discover (i) polymerase convoys, (ii) the multiscale nature of gene bursting, in which different time scales of promoter fluctuations are controlled by distinct promoter elements, and (iii) the stochastic nature or promoter proximal pausing (Tantale, Nat Comm 2016, Nat Comm 2021). These processes are directly relevant for HIV-1 latency.
Now, our main goal is to integrate biochemical and imaging data to provide a molecular understanding of promoter dynamics. Indeed, gene transcription levels are encoded in the rates controlling the intrinsic dynamic properties of promoters. Live cell transcription imaging can reveal these rates but cannot identify their molecular nature. To address this long-standing question, we are combining Single Molecule DNA Footprinting (SMF) and live cell transcription imaging. By doing so, we simultaneously capture the kinetic and molecular aspects of promoter fluctuations, and we can reveal the mechanisms of promoter dynamics. In parallel, we are developping tools to scale-up transcription imaging and perform live analyses in physiologically relevant models.
2- Spatio-temporal regulation of RNA localization and translation
Most mRNAs distribute randomly in the cytoplasm, but some localize to specific cellular sub-structures. Specific localizations have been observed from bacteria and yeast to metazoans, where they occur in oocytes, early embryos and somatic cells (see Bovaird, FEBS Letters 2018 for a review). RNA localization is linked to either RNA metabolism, e.g. for storage or decay; or to protein metabolism, to synthesize a protein locally. Such a local synthesis has been shown to play a role in many fundamental cellular processes, including cell polarity, mitosis and synaptic plasticity. However, we lacked a global view of RNA localization in human cells and this was the primary goal of this line of research.
We performed mRNA localization screens using single molecule FISH (smFISH). We screened thousands of mRNAs and found dozens that display specific cytoplasmic localizations. We observed local translation at unexpected locations, including cytoplasmic protrusions, cell edges, endosomes, the Golgi apparatus, the nuclear envelope and centrosomes (Chouaib et al., Dev Cell, 2020). We made two surprising observations. First, mRNA localization frequently required ongoing translation, indicating widespread co-translational targeting of mRNAs. This was a major paradigm shift, as it contradicts the current dogma in which the mRNA is not translated while transported. Second, we identified mRNAs that accumulated in cytoplasmic foci that were translation sites, i.e., specialized translation factories. We found that translation factories are polysome condensates and that their formation frequently involves multivalent interactions with the nascent protein chain. These discoveries profoundly changed our vision of the cellular organization of translation.
We recently studied the translation factories formed by β-catenin mRNA. β-catenin is a multi-functional protein playing essential roles in tissue homeostasis and cancer. It bridges E-cadherin to the cytoskeleton and also activates transcription of target genes in response to Wnt signaling. Plasma membrane β-catenin is stable whereas without Wnt, cytoplasmic β-catenin is degraded by the destruction complex, composed of APC and Axin. We found that the β-catenin translation factories accumulated APC and Axin and were dissolved by Wnt signaling. This indicated a model where translation factories were involved in the degradation of the nascent protein, thus uniquely regulating protein metabolism.
Now, we are studying how co-translational interactions control RNA localization, using centrosomal mRNAs as models. In particular, we aim at understanding how co-translational interactions are coupled to ribosome elongation and polysome transport. We are also interested in determining the function of this local translation.
3- Assembly of macromolecular complexes by the HSP90/R2TP chaperone
The assembly of macromolecular complexes is an essential cellular process. It often requires specialized factors that integrate together tertiary and quaternary folding. The HSP90/R2TP chaperone plays a unique role in the cell as a generalist assembly machine. This chaperone consists of a RPAP3:PIH1D1 hetero-dimer associated with the RUVBL1 and RUVBL2 AAA+ ATPases, themselves forming a hetero-hexamer. The HSP90/R2TP chaperone system is involved in the assembly of a many macromolecular complexes (Figure 1A), including non-coding RNPs (snRNPs, snoRNPs, miRNPs, the telomerase RNP), the three nuclear RNA polymerases and the kinases of the PIKK family. Overall, it is estimated that the R2TP chaperone is involved in the assembly of ~100 client complexes. The assembly mechanism used by R2TP remains incompletely characterized. The current model is that the hexameric nature of RUVBL1/RUVBL2 enables them to grab and bring together multiple subunits, with ATP regulating their binding and release, possibly by inducing large scale conformational changes (see our review Houry et al., TiBS 2018).
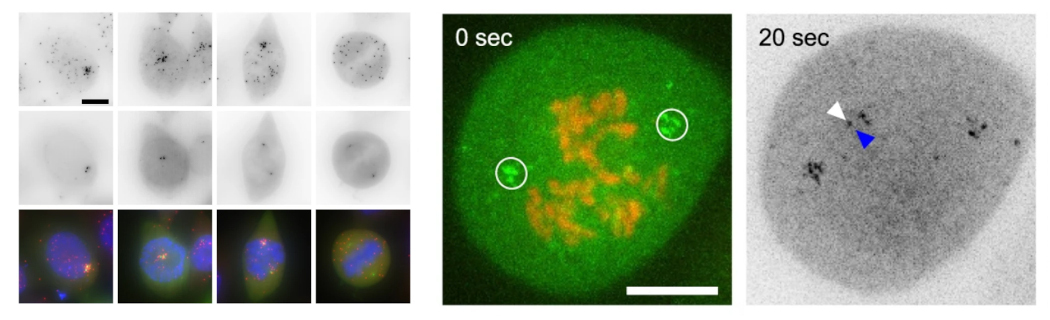
Our most recent interest focuses on the links between translation and the assembly of macromolecular complexes. In particular, we are interested in determining whether cells have ‘assembly lines’, which would start with protein synthesis and terminate with the delivery of a fully assembled, functional machinery. In agreement, we have found that R2TP binds co-translationally to many of its partners. We discovered that the HSP90 and R2TP chaperones induce the formation of translation factories that concentrate their client mRNAs and favour co-translational interactions with the chaperones (Figure 1B). It is remarkable that chaperones drive the compartmentalization of translation in the cytoplasm.
Now, we want to understand how co-translational interactions control the assembly of newly-synthesized proteins in macromolecular complexes. We aim to address how ribosome elongation is regulated and whether it is linked to the formation of the R2TP translation factories. We also would like to determine the assembly steps occurring in these factories. Another equally important aspect is to clarify the role of the HSP90/R2TP in cancer. Numerous pieces of evidence link the activity of this chaperone to tumorigenesis, but the mechanisms, and how they can be harnessed to fight cancer, remain poorly understood. Small molecules targeting R2TP have been recently developped and they show anti-tumor effects in mice. We aim at using these drugs to better understand the function of R2TP in tumorigenesis, in particular for a type of colorectal tumors that is difficult to treat.






























- Chemical inhibition of Exon Junction Complex assembly impairs mRNA localization and neural stem cells ciliogenesis. Nucleic Acids Res., in press Villa, T., Pourcelot O., Dierks, D., Faucourt, M., Slimani, F., Burel, C., Guyonnet, L., Spassky, N., Schwartz, S., Bertrand, E., Bensaude, O., Le Hir H.*
- Uniform dynamics of cohesin-mediated loop extrusion. Nature Genetics, in press. Sabaté, T., Lelandais, B., Robert, MC., Szalay, M., Tinevez, JY., Bertrand, E.*, Zimmer, C.*.
- The purification of biomolecular condensates – bottlenecks and strategies. J. Cell Biol., in press. Tartier,S., Basbous, J., Boulon, S., Verheggen, C., and E. Bertrand*
- eIF2A regulates cell migration in a translation-independent manner. Science Advances, in press. Jungfleisch, J., Mestre-Farràs, N., Pourcelot, O., Gómez-Riera, R., Halidi, N., Bertrand, E., and F. Gebauer*
- Polysome sorting: a mechanism for mRNA localization and local translation. Trends in Cell Biol. in press. Safieddine, A., Bizarro, J., Salloum, S., Le Hir H. and E. Bertrand*
- G-quadruplexes are promoter elements controlling nucleosome exclusion and RNA polymerase II pausing. Nat Genetics, 2025, 57:1981-1993. Esnault, C., Garcia-Oliver, E., Zine El Aabidine, A., Robert, MC., Magat, T., Gawron, K., Basyuk, E., Karpinska, M., Pigeot, A., Cucchiarini, A., Luo, Y., Verga, D., Mourad, R., Radulescu, O., Mergny, JL., Bertrand, E., and JC. Andrau*
- Identifying Markov chain models from time-to-event data: an algebraic approach. Bull Math Biol. 2024, 87:11. Radulescu*, O., Grigoriev, D., Seiss, M., Douaihy, M., Lagha, M., Bertrand, E. doi: 10.1007/s11538-024-01385-y.
- The HSP90/R2TP quaternary chaperone scaffolds assembly of the TSC complex. J. Mol Biol, 2024, 436:168840. Abéza, C., Chagot, M-E., Schneider, J., Robert, MC., Vandermoere, F., Schaeffer, C., Charpentier, B., Manival, X., Cianferani, X., Bertrand, E.* and C. Verheggen*
- Boosting the toolbox for live imaging of translation. RNA. 2024, rna.080140.124. Bellec, M., Chen, R., Dhayni, J., Trullo, A., Avinens, D., Karaki, H., Mazzarda, F., Lenden, H., Favard, C., Lehmann, R., Bertrand, E.*, Lagha, M. and J. Dufourt*
- The nucleolar phase of signal recognition particle assembly. Life Sci Alliance. 2024,7:e202402614. Issa, A., Schlotter, F., Flayac, J., Chen, J., Wacheul, L., Philippe, M., Sardini, L., Mostefa, L., Vandermoere, F., Bertrand, E., Verheggen, C., Lafontaine, DL., and S. Massenet*
- smFISH for Plants. Methods Mol Biol. 2024; 2784:87-100. Hani, S., Mercier, C., David, P., Desnos, T., Escudier, JM., Bertrand, E. and L. Nussaume*
- Cell cycle-dependent mRNA localization in P-bodies. Mol. Cell, 2024, 84:4191-4208. Safieddine, A., Benassy, MN., Bonte, T., Slimani, F., Pourcelot, O., Kress, M., Ernoult-Lange, M., Courel, M., Coleno, E., Imbert, A., Laine, A., Munier Godebert, A., Vinit, A., Blugeon, C., Chevreux, G., Gautheret, D., Walter, T., Bertrand, E., Bénard, M., and D. Weil.
- Transcriptional stochasticity as a key aspect of HIV-1 latency. Viruses 2023, 15:1969. Damour, A., Slaninova, V., Radulescu, O., Bertrand, E.*, and E. Basyuk*. Invited review
- Chromatin organization drives the search mechanism of nuclear factors. Nat Comm. 2023 14:6433. Mazzocca, M., Loffreda, A., Colombo, E., Fillot, T., Gnani, D., Falletta, P., Monteleone, E., Capozi, S., Bertrand E, Legube, G., Lavagnino, Z., Tacchetti, C. and Mazza D.
- Learning with Minimal Effort: Leveraging in Silico Labeling for Cell and Nucleus Segmentation 2023, Computer Vision – ECCV 2022 Workshops – Chapter Bonte, T., Philbert, M., Coleno, E., Bertrand, E., Imbert, A., and T. Walter.
- Proteomic analyses reveal new features of the box H/ACA RNP biogenesis. Nucleic Acids Res. 2023, 51:3357-3374. Schlotter, F., Mérouani, S., Flayac, J., Kogey, V., Issa, A., Dodré, M., Huttin, A., Branlant, C., Bertrand, E., Labialle, S., Vandermoere, F., Verheggen, C., Massenet S.
- Changes in adenoviral chromatin organization precede early gene activation upon infection. EMBO J. 2023 42:e114162. Schwartz, U., Komatsu, T., Huber, C., Lagadec, F., Baumgartl, C., Silberhorn, E., Nuetzel, M., Rayne, F., Basyuk, E., Bertrand, E., Rehli, M., Wodrich, H. and Laengst, G.
- Polymer simulations guide the detection and quantification of chromatin loop extrusion by imaging. Nucleic Acids Res., 2023, 51:2614-2632. Sabaté, T., Lelandais, B., Bertrand, E.*, Zimmer, C.*
- BurstDECONV: a signal deconvolution method to uncover mechanisms of transcriptional bursting in live cells. Nucl. Acids Res., 2023, 51:e88. Douathy, M.+, Topno, R.+, Lagha, M., Bertrand, E.*, Radulescu, O.* +: equal contribution
- HT-smFISH: a cost-effective and flexible workflow for high-throughput single-molecule RNA imaging. Nature Protocol, 2023, 18:157-187. Safieddine, A., Coleno, E., Lionneton, E., Traboulsi, A-M., Salloum, S., Lecellier, C-H., Gostan, T., Georget, V., Hassen-Khodja, C., Imbert, A., Müeller, F., Walter, T., Peter, M., Bertrand, E.
- Deciphering cellular and molecular determinants of human DPCD protein in complex with RUVBL1/RUVBL2 AAA-ATPases. J Mol Biol. 2022, 434:167760. Dos Santos Morais, R., Santo, PE., Ley, M., Schelcher, C., Abel, Y., Plassart, L., Deslignière, E., Chagot, ME., Quinternet, M., Paiva, ACF., Hessmann, S., Morellet, N.M.F., Sousa, P., Vandermoere, F., Bertrand, E., Charpentier, B., Bandeiras, TM., Plisson-Chastang, C., Verheggen, C., Cianférani, S., Manival, X.
- Long non-coding RNA Neat1 and paraspeckle components are translational regulators in hypoxia.
- Elife. 2022,11:e69162. Godet, AC., Roussel, E., David, F., Hantelys, F., Morfoisse, F., Alves, J., Pujol, F., Ader, I., Bertrand, E., Burlet-Schiltz, O., Froment, C., Henras, AK., Vitali, P., Lacazette, E., Tatin, F., Garmy-Susini, B. and Prats, AC.
- Versatile CRISPR-Based Method for Site-Specific Insertion of Repeat Arrays to Visualize Chromatin Loci in Living Cells. Methods Mol Biol. 2022;2532:275-290. Sabate, T., Zimmer, C., Bertrand, E.
- FISH-quant v2: a scalable and modular tool for smFISH image analysis 2022, RNA. 2022, 2:786-795. Imbert, A., Ouyang, W., Safieddine, A., Coleno, E., Zimmer, C., Bertrand, E., Walter, T., and Mueller, F.
- The interaction between RPAP3 and TRBP reveals a possible involvement of the HSP90/R2TP chaperone complex in the regulation of miRNA activity 2022, Nucleic Acids Res., 50:2172-2189. Abel, Y., Charron, C., Virciglio, C., Bourguignon-Igel, V., Quinternet, M., Chagot, ME., Robert, MC., Verheggen, C., Branlant, C., Bertrand, E., Manival, X., Charpentier, B., and M. Rederstorff.
- GOLT1B Activation in Hepatitis C Virus-Infected Hepatocytes Links ER Trafficking and Viral Replication 2021, Pathogens, 11:46. Butterworth, J., Gregoire, D., Peter, M., Roca Suarez, A.A., Desandré, G., Simonin, Y., Virzì, A., Zine El Aabidine, A., Guivarch, M., Andrau, JC., Bertrand, E., Assenat, E., Lupberger, J., and U. Hibner.
- The kinesin KIF1C transports AP-dependent mRNAs to cell protrusions 2021, RNA, 27:1528-1544. Pichon, X., Moissoglu, K., Coleno, E., Wang, T., Imbert, A., Robert, MC., Peter, M., Chouaib, R., Walter, T., Mueller, F., Zibara, K., Bertrand, E.* and S. Mili*
- RNA labeling technology grants access to live plant single cell transcriptional dynamics: application to phosphate repression signaling cascade 2021, Nature Plants, 7, 1750-64. Hani, S., Laura Cuyas, L., David, P., Secco, D., Whelan, J., Thibaud, MC., Müller, F., Pochon, N., Javot, H., Merret, R., Faklaris, O., Maréchal, E., Bertrand, E.*, and L. Nussaume*.
- Live-cell imaging reveals the spatiotemporal organization of endogenous RNA polymerase II phosphorylation at a single gene 2021, Nature Comm, 12:3158. Forero-Quintero, L. S., Raymond, W., Handa, T., Saxton, M., Morisaki, T., Kimura, H., Bertrand, E., Munsky, B. and T. J. Stasevich
- Non-canonical argonaute loading of extracellular vesicle-derived exogenous single-stranded miRNA in recipient cells 2021, J. Cell Sci. 134:jcs253914. Ghoshal, B., Bertrand, E. and SN Bhattacharyya
- The box C/D snoRNP assembly factor Bcd1 interacts with the histone chaperone Rtt106 and controls its transcription dependent activity 2021, Nat. Comm, 12:1859. Bragantini, B., Charron, C., Bourguet, M., Paul, A., Tiotiu, D., Rothé, B., Marty, H., Terral, G., Hessmann, S., Decourty, L., Chagot, ME., Strub, JM., Massenet, S., Bertrand, E., Quinternet, M., Saveanu, C., Cianférani, S., Labialle, S., Manival, X. and B. Charpentier.
- Exon Junction Complex dependent mRNA localization is linked to centrosome organization during ciliogenesis 2021, Nat. Comm, 12:1351. Kwon, OH., Mishra, R., Safieddine, A., Coleno, E., Alasseur, Q., Faucourt, M., Barbosa, I., Bertrand, E., Spassky, N. and Le Hir, H. Preprint available at BioRxiv 2020.10.28.358960; doi: https://doi.org/10.1101/2020.10.28.358960.
- A choreography of mRNAs at centrosomes reveals a localization mechanism involving active polysome transport. 2021, Nat. Comm, 12:1352. Safieddine, A., Coleno, E., Traboulsi, A., Kwon OS., Lionneton, F., Georget, V., Robert, MC., Gostan, T., Lecellier, C., Salloum, S., Chouaib, R., Pichon, X., Le Hir, H. , Zibara, K., Peter, M., and E. Bertrand. Preprint available at BioRxiv 2020.09.04.282038; doi: https://doi.org/10.1101/2020.09.04.282038
- Stochastic pausing at latent HIV-1 promoters generates transcriptional bursting. 2021, Nat. Comm, 12:4503. Tantale, K., Garcia-Oliver, E., L’Hostis, A., Yang, Y., Robert, MC., Gostan, T., Basu, M., Kozulic-Pirher, A., Andrau, JC., Muller, F., Basyuk, E.*, Radulescu, O.*, and E. Bertrand*
- Quantitative imaging of transcription in living Drosophila embryos reveals the impact of core promoter motifs on promoter state dynamics. Nat Comm, 2021, 12:4504. Pimmet, V., Dejean, M., Fernandez, C., Trullo, A., Bertrand, E., Radulescu, O., Lagha, M.
- The R2TP chaperone assembles cellular machineries in intestinal CBC stem cells and progenitors. Nat. Comm, 2021, 12:4810. Maurizy, C., Abeza, C., Pinet, V., Ferrand, M., Paul, C., Bremond, J., Langa, F., Gerbe, F., Jay, P., Verheggen, C., Tinari, N., Helmlinger, D., Lattanzio, R., Bertrand, E.*, Hahne, M.*, and B. Pradet-Balade*
- NOPCHAP1 is a PAQosome cofactor that helps loading NOP58 on RUVBL1/2 during box C/D snoRNP biogenesis. Nucleic Acids Res, 2021, 49:1094:1113. Abel, Y., Paiva, A. C., Bizarro, J., Chagot, M.-E., Santo, P. E., Robert, M.-C., Quinternet, M., Vendermoere, F., Sousa, P. M., Fort, P., Charpentier, B., Manival, X., Bandeiras, T. M., Bertrand, E.*, and Verheggen, C*
- RNA transport from transcription to localized translation: a single molecule perspective. RNA Biol. 2020. doi: 10.1080/15476286.2020.1842631. Basyuk, E., Rage, F., and E. Bertrand.
- NF-kB triggers a prompt and sharp transcriptional response in cell populations that emerges from heterogeneous bursting in single cells. iScience, 2020, 23:101529. Zambrano, S., Loffreda, A., Carelli, E., Stefanelli, G., Colombo, F., Bertrand, E., Tacchetti, C., Agresti, A., Bianchi, M. E., Molina,, N., and D. Mazza.
- A localization screen reveals compartmentalized translation and widespread co-translational protein targeting. Developmental Cell, 2020, 54:773-791. Chouaib, R., Safieddine, A., Pichon, X., Kwon, OS., Samacoits, A., Traboulsi, AM., Tsanov, N., Robert, MC., Coleno, E., Poser, I., Zimmer, C., Hyman, A. A., Le Hir, H., Zibara, K., Peter, M., Mueller, F.*, Walter, T.*, and E. Bertrand*
- New generations of MS2 variants and MCP fusions to detect single mRNAs in living eukaryotic cells. Methods in Mol. Biol., 2020, 2166:121-144. Pichon, X., Robert, MC., Bertrand, E., Singer, RH., Tutucci, E.
- Live cell imaging reveals 3′-UTR dependent mRNA sorting to synapses. Nat. Comm, 2019, 10:3178. Bauer, K., Segura, I., Gaspar, I., Scheuss, V., Illig, C., Ammer, G., Hutten, S., Basyuk, E., Fernández-Moya, SM., Ehses, J., Bertrand, E., Kiebler, MA.
- A deep learning approach to identify mRNA localization patterns. Proceedings of the 16th IEEE International Symposium on Biomedical Imaging (ISBI 2019), accepted for publication. R. Dubois, A. Imbert, A. Samacoits, M. Peter, E. Bertrand, F. Müller & T. Walter.
- A computational framework to study sub-cellular mRNA localization. Nat. Comm, 2018, 9:4584. Samacoits, A., Chouaib, R., Safieddine, A., Traboulsi, A., Ouyang, W., Zimmer, C., Peter, M., Bertrand, E.*, Walter, T.*, Mueller, F.*
- The splicing factor SRSF3 is functionally connected to the nuclear RNA exosome for intronless mRNA decay. Sci Rep., 2018, 8:12901. Mure F, Corbin A, Benbahouche NEH, Bertrand E, Manet E, Gruffat H.
- A growing toolbox to image gene expression in single cells: sensitive approaches for demanding challenges. Mol. Cell, 2018, 71:468-480. Pichon, X., Lagha, M., Mueller, F. and Bertrand, E.
- The Role of Supercoiling in the Motor Activity of RNA Polymerases. Methods Mol Biol. 2018, 1805:215-232. Lesne A, Victor JM, Bertrand E, Basyuk E, Barbi M.
- Deep structural analysis of RPAP3 and PIH1D1, two components of the HSP90 co-chaperone R2TP complex. Structure, 2018, 26:1196-1209. Henri, J., Chagot, ME., Bourguet, M., Abel, Y., Terral, G., Maurizy, C., Aigueperse, C., Georgescauld, F., Vandermoere, F., Saint-Fort, R., Behm-Ansmant, I., Charpentier, B., Pradet-Balade, B., Verheggen, C., Bertrand, E., Meyer, P., Cianférani, S., Manival, X., and Quinternet, M.
- The RPAP3-Cterminal domain identifies R2TP-like quaternary chaperones. Nat Comm, 2018, 9:2093. Maurizy, C., Quinternet, M., Abel, Y., Verheggen, C., Santo, P. E., Bourguet, M., Paiva, A. C. F., Bragantini, B., Chagot, ME., Robert, MC., Abeza, C., Fabra, P., Fort, P., Vandermoere, F., Sousa, P., Rain, JC., Charpentier, B., Cianférani, S., Bandeiras, T. M., Pradet-Balade, B., Manival, X., Bertrand, E.
- Meg3 non-coding RNA expression controls imprinting by preventing transcriptional upregulation in cis. Cell Rep. 2018, 23:337-348. Sanli I, Lalevée S, Cammisa M, Perrin A, Rage F, Llères D, Riccio A, Bertrand E, Feil R.
- The PAQosome, an R2TP-based chaperone for quaternary structure formation. TIBS 2018, 43:4-9. Houry, W.A.*, Bertrand, E.*, and Coulombe, B.* *:co-corresponding authors.
- P-Body Purification Reveals the Condensation of Repressed mRNA Regulons. Mol. Cell, 2017, 68:1-14, Hubstenberger, A., Courel ,M., Bénard, M., Souquere, S., Ernoult-Lange, M., Chouaib, R., Yi, Z., Morlot, JB., Munier, A., Fradet M., Daunesse, M., Bertrand, E., Pierron, G., Mozziconacci, J., Kress, M., Weil, D.
- ARS2 is a general suppressor of pervasive transcription. Nucl. Acids Res., 2017, 45:10229-10241. Iasillo, C., Schmid, M., Yahia, Y., Maqbool, M., Descotes, N., Karadoulama, E., Bertrand, E., Andrau, JC., Jensen, T.
- Assembly of the U5 snRNP component PRPF8 is controlled by the HSP90/R2TP chaperones. J. Cell Biol., 2017, 21:1579-1596. Malinová, A., Cvačková, Z., Matějů, D., Hořejší, Z., Abéza, C., Vandermoere, F., Bertrand, E.*, Staněk, D.*, Verheggen, C.* *. co-corresponding authors.
- Mutually exclusive CBC-containing complexes contribute to RNA fate. Cell Reports, 2017, 18:2635-2650. Giacometti, S., Benbahouche, N. H., Domanski, M., Robert, M-C., Meola, N., Lubas, M., Bukenborg, J., Andersen, J. S., Schultze, W. M., Verheggen, C., Kudla, G.*, Jensen, T. H.*, Bertrand, E.*
- Assembly and trafficking of box C/D and H/ACA snoRNPs, RNA biology, 2017, 14:680-692. Massenet S., Bertrand E., Verheggen C.
- Visualization of single polysomes reveals translation dynamics of endogenous mRNAs in living human cells, J. Cell Biol., 2016, 214:769-81. Pichon X., Bastide A., Safieddine A., Chouaib R., Samacoits A., Basyuk E., Peter M., Mueller F., Bertrand E.
- SmiFISH and FISH-quant – a flexible single mRNA detection approach with super-resolution capability. Nucleic Acids Res, 2016, pii:gwk784, Tsanov, N., Samacoits, A., Chouaib, R., Traboulsi, A.M., Gostan, T., Weber, C., Zimmer, C., Zibara, K., Walter, T., Peter, M.*, Bertrand, E.*, Mueller, F* *. co-corresponding authors.
- Imaging HIV-1 RNA dimerization in cells by multicolor super-resolution and fluctuation microscopies. Nucleic Acids Res., 2016, 44:7922-34. Ferrer M, Clerté C, Chamontin C, Basyuk E, Lainé S, Hottin J, Bertrand E, Margeat E, Mougel M.
- A real-time, single molecule view of transcription reveals convoys of RNA polymerases and multiscale bursting. Nat. Comm., 2016, 7:12248. Tantale, K., Müller, F., Kozulic-Pirher, A., Lesne, A., Victor, JL., Robert, MC., Capozi, S., Bäcker, V., Mateos-Langerak, J., Darzacq, X., Zimmer, C., Basyuk, E., Bertrand, E.
- The in vivo dynamics of TCERG1, a factor that couples transcriptional elongation with splicing. RNA. 2016, 22:571-82Sánchez-Hernández N., Boireau S., Schmidt U., Muñoz-Cobo JP, Hernández-Munain C, Bertrand E., Suñé C.
- SnoRNPs, ZNHIT proteins and the R2TP pathway. Oncotarget. 2015, 6:41399-400. Verheggen, C., Pradet-Balade, B., Bertrand E. 2015, 6:41399-400
- NUFIP and the HSP90/R2TP chaperone bind the SMN complex and facilitate assembly of U4-specific proteins. Nucl Acids Res, 2015, 43:8973-87. Bizarro, J., Dodré, M., Huttin, A., Charpentier, B., Schlotter, F., Branlant, C., Verheggen, C., Massenet, S., Bertrand, E.
- Long lasting control of viral rebound with a new drug ABX464 targeting Rev-mediated viral RNA biogenesis. Retrovirology. 2015, 12:30. Campos N, Myburgh R, Garcel A, Vautrin A, Lapasset L, Nadal ES, Mahuteau-Betzer F, Najman R, Fornarelli P, Tantale K, Basyuk E, Séveno M, Venables JP, Pau B, Bertrand E, Wainberg MA, Speck RF, Scherrer D, Tazi J.
- Proteomic and 3D structure analyses highlight the C/D box snoRNP assembly mechanism and its control. J. Cell Biol., 2014; 207:463-480. Bizarro, J., Charron, C., Boulon, S., Westman, B., Pradet-Balade, B., Vandermoere, F., Chagot, ME., Hallais, M., Ahmad, Y., Leonhardt, H., Lamond, A., Manival, X., Branlant, C., Charpentier, B., Verheggen*, C., Bertrand*, E.
- Hypermethylated-capped selenoprotein mRNAs in mammals. Nucleic Acids Res. 2014;42(13):8663-77. Wurth L, Gribling-Burrer AS, Verheggen C, Leichter M, Takeuchi A, Baudrey S, Martin F, Krol A, Bertrand E, Allmang C.
- MLN51 triggers P-body disassembly and formation of a new type of RNA granules. J Cell Sci. 2014, 127:4692-701. Cougot N, Daguenet E, Baguet A, Cavalier A, Thomas D, Bellaud P, Fautrel A, Godey F, Bertrand E, Tomasetto C, Gillet R.
- Characterization of spaghetti function in Drosophila supports a role for Hsp70 in R2TP/Hsp90-assisted assembly of cellular machineries. J Biol Chem, 2014, 289(9):6236-47. Benbahouche, H., Iliopoulos, I., Török, I., Marhold, J., Kajava, A., Kempf, T., Schnölzer, M., Kiss, I., Bertrand, E.*, Mechler*, B. M.*, Pradet-Balade, B.*
- Stable assembly of HIV-1 export complexes occurs co-transcriptionally. RNA, 2014, 20:1-8. Nawroth, I., Mueller, F., Basyuk, E., Beerens, N., Rahbek, U. Darzacq, X., Bertrand, E.*, Kjems, J.* and Schmidt, U.*
- Identification of the interface between Snu13p/15.5K and Rsa1p/NUFIP and demonstration of its functional importance for snoRNP assembly. Nucl Acids Res, 2014, 42(3):2015-36. Rothé, B., Back, R., Quinternet, M., Bizarro, J., Robert, M-C., Blaud, M., Romier, C., Manival, X.*, Charpentier, B.*, Bertrand, E.*, Branlant, C.
- CBC-ARS2 stimulate 3′-end maturation of multiple RNA families and favor cap-proximal processing. Nat Struct Mol Biol, 2013, 20:1367-76. Hallais, M., Pontvianne, F., Refsing Andersen, P., Clerici, M., Lener, D., Benbahouche, H., Gostan, T.,Vandermoere, F., Robert, M-C., Cusack, S., Verheggen, C., Jensen, T. H. and Bertrand, E.
- The human cap-binding complex is functionally connected to the nuclear RNA exosome. Nat Struct Mol Biol, 2013, 20:1358-66. Refsing Andersen, P., Domanski, M., Kristiansen, M., Storvall, E., Ntini, E., Verheggen, C., Bunkenborg, J., Poser, I., Hallais, M., Sandberg, R., Hyman, A., LaCava, J., Rout, M. P., Andersen, J. S., Bertrand, E., and Jensen, T. H.
- Genome-wide identification of mRNAs associated with the Survival of Motor. RNA, 2013, 19:1755-66. Rage, F., Boulisfane, N., Rihan, K., Neel, H., Gostan, T., Bertrand, E., Bordonné, R., and Soret, J.
- FISH-quant : automated counting of transcripts in 3D images. Nat Methods. 2013, 10:277-8. Mueller, F., Senecal, A., Tantale, K., Marie-Nelly, H., Ly, N., Collin, O., Basyuk, E., Bertrand, E.*, Darzacq, X.* and Zimmer, C*.
- Nuclear retention prevents premature cytoplasmic appearance of mRNA. Mol Cell, 2012, 48:145-152. Kallehauge,T., Robert, M-C., Bertrand*, E., Jensen*, T. H.
- CRM1 plays a nuclear role in transporting snoRNPs to nucleoli in higher eukaryotes. Nucleus. 2012. 3(2). Verheggen C, Bertrand E.
- Microprocessor dynamics and interactions at endogenous imprinted C19MC microRNA genes. J Cell Sci. 2012, 125:2709-20. Bellemer C, Bortolin-Cavaillé ML, Schmidt U, Jensen SM, Kjems J, Bertrand E, Cavaillé J.
- Perispeckles are major assembly sites for the exon junction core complex. Mol Biol Cell. 2012. 23(9):1765-82. Daguenet E, Baguet A, Degot S, Schmidt U, Alpy F, Wendling C, Spiegelhalter C, Kessler P, Rio MC, Le Hir H, Bertrand E, Tomasetto C.
- HSP90 and the R2TP co-chaperone complex: building multi-protein machineries essential for cell growth and gene expression. RNA Biol, 2012, 9: 148-54. Boulon, S., Bertrand, E.*, and B. Pradet-Balade*.
- Retroviral GAG proteins recruit AGO2 on viral RNAs without affecting RNA accumulation and translation (2012). Nucleic Acids Res. 40, 775-786. Bouttier M., Saumet A., Peter M., Courgnaud V., Schmidt U., Cazevieille C., Bertrand E., Lecellier CH.
- CRM1 controls the composition of nucleoplasmic pre-snoRNA complexes to licence them for nucleolar transport. EMBO J., 2011, 30:2205-2218. Pradet-Balade, B., Girard, C., Boulon, S., Paul, C., Azzag, K., Bordonne, R., Bertrand, E.*, and Verheggen, C*
- Real-time imaging of co-transcriptional splicing reveals a kinetic model that reduces noise: implication for alternative splicing regulation. J Cell Biol, 2011, 193:819-829. Schmidt, U., Basyuk, E., Robert, MC., Yoshida, M., Villemin, JP., Auboeuf, D., Aitken, S. and Bertrand, E.
- Real-time imaging of the HIV-1 transcription cycle in living cells. Methods. 2011, 53:52-57.Maiuri P, Knezevich A, Bertrand E, Marcello A.
- Crosstalk between mRNA 3′-end processing and transcription initiation. Mol Cell, 2010, 40:410-422. Mapendano, C., Lykke-Andersen, S., Kjems, J., Bertrand, E. and T. H. Jensen.
- HSP90 and its R2TP/Prefoldin-like co-chaperone are involved in the cytoplasmic assembly of RNA polymerase II. Mol Cell, 2010, 39:912-24. Boulon, S., Pradet-Balade, B., Verheggen, C., Molle, D., Boireau, S., Georgieva, M., Azzag, K., Robert, M-C., Ahmad, Y., Neel, H., Lamond, A.I., Bertrand, E.
- RiboSys, a high-resolution, quantitative approach to measure the in vivo kinetics of pre-mRNA splicing and 3′-end formation processing in S. Cerevisiae. RNA, 2010.16(12):2570-80. Alexander RD, Barrass JD, Dichtl B, Kos M, Obtulowicz T, Robert MC, Koper M, Karkusiewicz I, Mariconti L, Tollervey D, Dichtl B, Kufel J, Bertrand E, Beggs JD.
- Splicing independent recruitment of U1 snRNA to a transcription unit in living cells. J Cell Sci. 2010, 123:2085-93. Spiluttini B, Gu B, Belagal P, Smirnova AS, Nguyen VT, Hébert C, Schmidt U, Bertrand E, Darzacq X, Bensaude O.
- A Proteomic Screen for Nucleolar SUMO Targets shows SUMOylation modulates the function of Nop5/Nop58. Mol Cell, 2010, 39:618-31. Westman, B.J., Verheggen, C., Hutten, S.,Lam, Y. W., Bertrand, E., Lamond, A. I.
- Establishment of a protein frequency library and its application in the reliable identification of specific protein interaction partners. Mol Cell Proteomics, 2010, 9:861-79.Boulon S, Ahmad Y, Trinkle-Mulcahy L, Verheggen C, Cobley A, Gregor P, Bertrand E, Whitehorn M, Lamond AI.
- Processivity and coupling in messenger RNA transcription. Plos One, 2010, 5:e8845. Aitken, S., Robert, M-C., Ross D. Alexander, Igor Goryanin, E.Bertrand, Jean D. Beggs.
- Assembly of an export-competent mRNP is needed for efficient release of the 3′-end processing complex after polyadenylation. Mol Cell Biol. 2009, 19:5327-38. Qu X, Lykke-Andersen S, Nasser T, Saguez C, Bertrand E, Jensen TH, Moore C.
- Endosomal trafficking of HIV-1 Gag and genomic RNAs regulates viral egress. J. Biol Chem, 2009, 284:19727-43. Molle, D., Segura-Morales, C., Camus, G., Berlioz-Torrent, C., Kjems, J., Basyuk, E., Bertrand, E.
- DNA Damage Regulates Alternative Splicing through Changes in Pol II Elongation. Cell, 2009, 137:708-20. Muñoz, M. J., Pérez Santangelo, S. M., de la Mata, M., Bird, G., Bentley, D., Boireau, S., Bertrand, E., Kornblihtt, A. R.
- Dendrites of mammalian neurons contain specialized P-Body-Like Structures that respond to neuronal activation. J. Neurosci. 2008, 28:13793-13804. Cougot N, Bhattacharyya SN, Tapia-Arancibia L, Bordonné R, Filipowicz W, Bertrand E, Rage F.
- Translationally repressed mRNA transiently cycles through stress granules during stress. Mol Biol Cell. 2008, 19(10):4469-79. Mollet S, Cougot N, Wilczynska A, Dautry F, Kress M, Bertrand E, Weil D.
- Mutations in a small region of the exportin Crm1p disrupt the daughter cell specific nuclear localization of the transcription factor Ace2p in Saccharomyces cerevisiae. Biol Cell. 2008, 100(6):343-54. Bourens, M., Racki, W., Bécam, AM., Panozzo, C., Boulon, S., Bertrand, E., Herbert, CJ.
- The HSP90 chaperone controls the biogenesis of L7Ae RNPs through a conserved machinery. J. Cell Biol, 2008, 180(3):579-95. Boulon, S., Marmier-Gourrier, N., Wurth, L., Pradet-Balade, B., Verheggen, C., Jády, B., Rothé, B., Pescia, C., Robert, M-C., Kiss, T., Bardoni, B., Krol, A., Branlant, C., Allmang, C., Charpentier, B., and Bertrand, E.
- A novel role for PA28gamma-proteasome in nuclear speckle organization and SR protein trafficking. Mol Biol Cell. 2008, 19(4):1706-16. Baldin V, Militello M, Thomas Y, Doucet C, Fic W, Boireau S, Jariel-Encontre I, Piechaczyk M, Bertrand E, Tazi J, Coux O.
- Characterization of a short isoform of human Tgs1 hypermethylase associating with snoRNPs core proteins and produced by limited proteasome processing. J Biol. Chem, 2008, 283(4):2060-9. Girard, C., Verheggen, C., Vagner, C., Bertrand, E., and Bordonné, R.
- The transcriptional cycle of HIV-1 in real-time and live cells. J. Cell Biol. 2007, 179:291-304. Boireau, S., Maiuri, P., Basyuk, E., de la Mata, M., Knezevich, A., Pradet-Balade, B., Bäcker, V., Kornblihtt, A., Marcello, A., and Bertrand E.
- Inhibition of nonsense-mediated mRNA decay (NMD) by a new chemical molecule reveals the dynamic of NMD factors in P-bodies. J Cell Biol. 2007, 178:1145-60. Durand, S., Cougot, N., Mahuteau-Betzer, F., Nguyen, C.H., Grierson, D.S., Bertrand, E., Tazi, J., and Lejeune, F.
- The exon-junction-complex-component metastatic lymph node 51 functions in stress-granule assembly. J Cell Sci. 2007, 120:2774-84. Baguet, A., Degot, S., Cougot, N., Bertrand, E., Chenard, MP., Wendling, C., Kessler, P., Le Hir, H., Rio, M-C., and Tomasetto, C.
- A dynamic scaffold of pre-snoRNP factors facilitates human box C/D snoRNP assembly. Mol Cell Biol. 2007, 27:6782-93. McKeegan, K., Debieux, C., Boulon, S., Bertrand, E., and Watkins, NJ.
- The clathrin adaptor complex AP-1 binds HIV-1 and MLV Gag and facilitates their budding. Mol Biol Cell. 2007, 18:3193-203. Camus, G., Segura-Morales, C., Molle, D., Lopez-Verges, S., Begon-Pescia, C., Cazevieille, C., Schu, P., Bertrand, E., Berlioz-Torrent, C., and Basyuk, E.
- A real-time view of the TAR:Tat:P-TEFb complex at HIV-1 transcription sites. Retrovirology. 2007, 4:36. Molle, D., Maiuri, P., Boireau, S., Bertrand, E., Knezevich, A., Marcello, A., and Basyuk, E.
- Bsr, a novel nuclear-restricted RNA with mono-allelic expression. Mol Biol Cell, 2007, 18:2817-27. Royo, H., Basyuk, E., Marty, E., Marques, M., Bertrand, E., and Cavaille, J.
- Suv39H1 and HP1gamma are responsible for chromatin-mediated HIV-1 transcriptional silencing and post-integration latency. EMBO J. 2007, 26:424-35. Du Chene, I., Basyuk, E., Lin, Y., Triboulet, R., Knezevich, A., Chable-Bessia, C., Mettling, C., Baillat, V., Reynes, J., Corbeau, P., Bertrand, E., Marcello, A., Emiliani, S., Kiernan, R., and Benkirane, M.
- Depletion of SMN by RNA interference in HeLa cells induces defects in Cajal body formation. Nucleic Acids Res. 2006, 34:2925-32. Girard, C., Neel, H., Bertrand, E., and Bordonne, R.
- The exonuclease ISG20 mainly localizes in the nucleolus and the Cajal (Coiled) bodies and is associated with nuclear SMN protein-containing complexes. J Cell Biochem. 2006, 98:1320-33. Espert, L., Eldin, P., Gongora, C., Bayard, B., Harper, F., Chelbi-Alix, M., Bertrand, E., Degols, G., and Mechti, N.
- Cell cycle-dependent recruitment of telomerase RNA and Cajal bodies to human telomeres. Mol Biol Cell. 2006, 17:944-54. Jady, B., Richard, P., Bertrand, E., and Kiss, T.
- Photo-conversion of YFP proteins into CFP-like species during acceptor photo-bleaching FRET experiments. Nature Methods, 2005, 2:801. Valentin, G., Verheggen, C., Piolot, T., Neel, H., Coppey-Moisan, M., and Bertrand, E.
- The packaging signal of MLV is an integrated module that mediates intracellular transport of genomic RNAs. J. Mol. Biol., 2005, 354:330-9. Basyuk, E., Boulon, S., Pedersen, F. S., Bertrand, E., and Rasmussen, S. V.
- Inhibition of translational initiation by Let-7 microRNA in human cells. Science, 2005, 309:1573-6. Pillai, R., Battacharrya, S., Artus, C., Zoller, T., Cougot, N., Basyuk, E., Bertrand, E., Filipowicz, W.
- Tsg101 and Alix interact with MLV Gag and cooperate with Nedd4 ubiquitin-ligases during budding. J. Biol. Chem, 2005, 280:27004-12. Segura-morales, C., Pescia, C., Chatellard-Causse, C., Sadoul, R., Bertrand, E. and Basyuk, E.
- ADAR2-mediated editing of RNA substrates in the nucleolus is inhibited by C/D small nucleolar RNA. J. Cell Biol. 2005, 169:745-753. Vitali, P., Basyuk, E., Le Meur, E., Bertrand, E., Muscatelli, F., Cavaille, J., and Huttenhofer, A.
- PHAX and CRM1 are required sequentially to transport U3 snoRNA to nucleoli. Mol Cell, 2004, 16:777-787. Boulon, S., Verheggen, C., Jady, B., Girard, C., Pescia, C., Paul, C., Ospina, J., Kiss, T., Matera, A. G., Bordonné, R. and Bertrand, E.
- A co-transcriptional model for 3′-end processing of the Saccharomyces cerevisiae pre-ribosomal RNA precursor. RNA. 2004 Oct;10(10):1572-85. Henras AK, Bertrand E, Chanfreau G.
- Nuclear localization properties of a conserved protuberance in the Sm core complex. Exp Cell Res. 2004 299(1):199-208. Girard C, Mouaikel J, Neel H, Bertrand E, Bordonne R.
- Human box H/ACA pseudouridylation guide RNA machinery. Mol Cell Biol. 2004 Jul;24(13):5797-807. Kiss AM, Jady BE, Bertrand E, Kiss T.
- From silencing to gene expression: real-time analysis in single cells. Cell, 2004, 116(5):683-98. Janicki, S., Tsukamoto, T., Salghetti, S., Tansey, W., Sachidanandam, R., Prasanth, K., Ried, T., Shav-Tal, Y., Bertrand, E., Singer, R., and Spector, D.
- Human telomerase RNA and box H/ACA scaRNAs share a common Cajal body localization signal. J. Cell Biol, 2004, 164(5):647-52. Jady, B., Bertrand, E., and Kiss, T.
- Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucleic Acids Res. 2003, 31(22):6593-7. Basyuk, E., Suavet, F., Doglio, A., Bordonné, R, Bertrand E.
- Retroviral genomic RNAs are transported to the plasma membrane by endosomal vesicles. Dev. Cell, 2003, 5:161-174. Basyuk, E., Galli, T., Mougel, M., Blanchard, JM., Sitbon, M., and Bertrand E.
- A common sequence motif determines the Cajal body-specific localisation of box H/ACA scaRNAs EMBO J. 2003, 22:4283-93. Richard, P., Xavier Darzacq, X., Bertrand, E., Jády, B., Verheggen, C., and Kiss, T.
- Interaction between the small-nuclear-RNA cap hypermethylase and the spinal muscular atrophy protein, survival of motor neuron. EMBO Rep., 2003, 4(6):616-22. Mouaikel J, Narayanan U, Verheggen C, Matera AG, Bertrand E, Tazi J, Bordonne R.
- Modification of Sm small nuclear RNAs occurs in the nucleoplasmic Cajal body following import from the cytoplasm. EMBO J., 2003, 22(8):1878-88. Jady BE, Darzacq X, Tucker KE, Matera AG, Bertrand E, Kiss T.
- The RasGAP-associated endoribonuclease G3BP assembles stress granules. J Cell Biol. 2003, 160(6):823-31. Tourriere H, Chebli K, Zekri L, Courselaud B, Blanchard JM, Bertrand E, Tazi J.
- Exportin-5 mediates nuclear export of minihelix-containing RNAs. J. Biol Chem. 2003, 278(8):5505-8. Gwizdek C, Ossareh-Nazari B, Brownawell AM, Doglio A, Bertrand E, Macara IG, Dargemont C.
- Single mRNA molecules demonstrate probabilistic movement in living mammalian cells Current Biology, 2003, 13(2):161-7. D. Fusco, N. Accornéro, S. Shenoy, JM Blanchard, RH Singer and E. Bertrand.
- A Cajal body-specific pseudouridylation guide RNA is composed of two box H/ACA snoRNA-like domains. Nucleic Acids Res. 2002, 30(21):4643-9. Kiss AM, Jady BE, Darzacq X, Verheggen C, Bertrand E, Kiss T.
- An active precursor in assembly of yeast nuclear ribonuclease P. RNA. 2002; 8(10):1348-60. Srisawat C, Houser-Scott F, Bertrand E, Xiao S, Singer RH, Engelke DR.
- Cajal body-specific small nuclear RNAs: a novel class of 2’-o-methylation and pseudouridylation guide RNAs. EMBO J., 2002, 21:2746-56. Xavier Darzacq, Beáta E. Jády, Céline Verheggen, Arnold M. Kiss, Edouard Bertrand and Tamás Kiss
- Mammalian and yeast U3 snoRNPs are matured in specific and related nuclear compartments. EMBO J., 2002, 21: 2736-45. Verheggen, C., Lafontaine, D., Samarsky, S., Mouaikel, J., Blanchard, JM., Bordonné, R., and Bertrand E.
- Hypermethylation of the cap structure of both snRNAs and snoRNAs in yeast requires a conserved methyltransferase that locates in the nucleolus. Mol. Cell, 2002, 9:891-901. John Mouaikel, Céline Verheggen, Edouard Bertrand, Jamal Tazi and Rémy Bordonné.
- Box C/D small nucleolar RNA trafficking involves small nucleolar RNP proteins, nucleolar factors and a novel nuclear domain. EMBO J. 2001, 20:5480-5490. Verheggen, C., Mouaikel, J., Thiry, M., Blanchard, J-M., Tollervey, D., Bordonné, R., Lafontaine, D., and Bertrand, E.
- A Well-Connected and Conserved Nucleoplasmic Helicase is Required for Production of Box C/D and H/ACA snoRNAs and Localization of snoRNP Proteins. Mol. Cell Biol. 2001, 21:7731-46. King, T., Decatur, W., Bertrand, E., Maxwell, E.S. and Fournier, M.J.
- Terminal minihelix, a novel RNA motif that directs polymerase III transcripts to the cell cytoplasm. J Biol Chem. 2001, 276(28):25910-8. Bertrand E, Gwizdek C, Dargemont C, Lefebvre JC, Blanchard JM, Singer RH, Doglio A.
- A CBF5 mutation that disrupts nucleolar localization of early tRNA biosynthesis in yeast also suppresses tRNA gene-mediated transcriptional silencing. Proc. Natl. Acad. Sci. USA, 2000, 97:13108-13. Ann Kendall, Melissa W. Hull, Edouard Bertrand, Paul D.Good, Robert H.Singer, and David R.Engelke.
- Alcaline fixation drastically enhances the signal of in situ hybridization. Nucl. Acids Res., 2000, 28:E46. Basyuk, E., Bertrand, E., Journot, L.
- mRNA localization signals can enhance the intracellular effectiveness of hammerhead ribozymes. RNA, 1999, 5:1200-1209. Lee NS, Bertrand E, Rossi J.
- A snoRNA:ribozyme hybrid cleaves a nucleolar RNA target in vivo with near-perfect efficiency. Proc. Natl. Acad. Sci. USA, 1999, 96:6609-14. Samarsky D, Ferbeyre G, Bertrand E, Singer RH, Cedergren R, Fournier MJ.
- Localization of ASH1 mRNA particles in living yeast. Molecular Cell, 1998, 2:437-445. Bertrand E, Chartrand P, Shaefer M, Shenoy S, Singer RH, Long R.
- 3’-end modification of the adenoviral VAI gene affects its expression in human cells: consequences for the design of chimeric VAI RNA-ribozymes. Antisense Nucleic Acid Drug Dev 1998, 8:379-90. Barcellini-Couget S, Bertrand E, Singer RH, Lebfevre JC, Doglio A.
- Monitoring retroviral RNA dimerization in vivo via hammerhead ribozyme cleavage. J. Virology, 1998 Oct;72(10):8349-53. Pal BK, Scherer L, Zelby L, Bertrand E, Rossi JJ.
- Nucleolar localization of early tRNA processing. Genes & Dev 1998 Aug 15, 12:2463-2468. Bertrand E, Houser-Scott F, Kendall A, Singer RH, Engelke DR.
- The snoRNA box C/D motif directs nucleolar targeting and also couples snoRNA synthesis and localization. EMBO J 1998 Jul 1; 17:3747-3757. Samarsky DA, Fournier MJ, Singer RH, Bertrand E.
- The expression cassette determines the functional activity of ribozymes in mammalian cells by controlling their intra-cellular localization. RNA 1997 Jan, 3(1):75-88. Bertrand E, Castanotto D, Zhou C, Carbonelle C, Lee NS, Good P, Chaterjee S, Grange T, Pictet R, Kohn D, Engelke DR, Rossi JJ.
- Expression of small, therapeutic RNAs in human cell nuclei. Gene Ther. 1997 Jan, 4:45-54. Good P, Krikos AJ, Li S, Bertrand E, Lee N, Giver L, Ellington A, Zaia JA, Rossi JJ, Engelke DR.
- Polystyrene reverse-phase ion-pair chromatography of chimeric ribozymes. Anal Biochem. 1994, 216:83-88. Swiderski PM, Bertrand E, Kaplan BE.
- Facilitation of hammerhead ribozyme catalysis by the nucleocapsid protein of HIV-1 and the heterogeneous nuclear ribonucleoprotein A1. EMBO J. 1994, 13:2904-12. Bertrand E, Rossi, JJ.
- Can hammerhead ribozymes be efficient tools to inactivate gene function? Nucleic Acids Res. 1994, 22:293-300. Bertrand E, Pictet R, Grange T.
- Visualization of the in vivo interaction of a regulatory protein with RNA. Proc. Natl. Acad. Sci. USA 1993, 90:3496-3500. Bertrand, E., Fromont-Racine, M., Pictet, R., and Grange, T.
- A highly sensitive method for mapping the 5′ termini of mRNAs. Nucleic Acids Res. 1993, 21:1683-1684. Fromont-Racine, M., Bertrand, E., Pictet, R., and Grange, T.






