Research teams
Molecular bases of human diseases
Genome surveillance and stability

Cells have evolved genome surveillance mechanisms to preserve genome stability. These include the cellular responses to DNA damage (DDR) ) and DNA damage tolerance involving translesion DNA synthesis (TLS). The DDR can slow down or arrest cell proliferation in the presence of DNA lesions generated by endogenous and exogenous sources of DNA damage, and as such avoid the propagation of mutations which are drivers of genomic instability, a hallmark of cancer. TLS is instead employed to tolerate unrepairable and/or excess DNA damage, thus avoiding cell death, and can be exploited by cancer cells to adapt to chemotherapeutic treatments. Hence, there is currently a great interest in targeting TLS to sensitize cancer cells resistant to therapy by identifying key novel TLS druggable targets.
To tackle these questions, we employ in vitro systems derived from fertilized eggs of the clawed frog Xenopus laevis, that also provide a performant model system for vertebrate development, as well as mouse and human pluripotent stem cells, cancer cell lines and 3D ex-vivo systems (tumoroids).
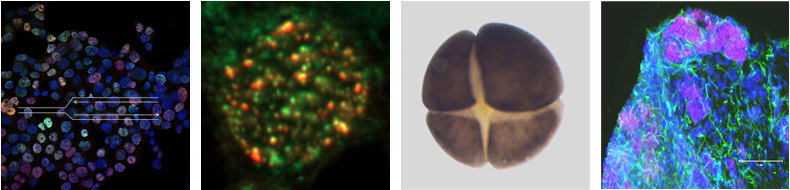
Identification of novel TLS factors
Antoine Aze, Quentin Debelley
TLS in initiated by monoubiquitination of the replication factor PCNA (PCNAmUb) by the Rad6(E2)/Rad18(E3) ubiquitin ligase complex. PCNAmUb triggers recruitment of Y-family TLS DNA polymerases (pols; Polƞ, k, ɩ and Rev1) into distinct nuclear foci whose molecular composition is currently not well defined. These structures, that we have called “the translesiosomes”, then perform the DNA lesion bypass, although at the expenses of increased mutagenesis, since TLS pols are error-prone (Maiorano D et al. Int J Mol Sci. 2021, for review). We have started to characterize the macromolecular composition of translesiosomes by a proteomic approach using Xenopus egg extracts that reproduce TLS activation (Aze et al., Methods in Molecular Biology, 2024). We have identified several novel interactors whose functional characterization is under way.
Characterization of TLS dynamics in single cells
James Hutchins
Translesiosomes have been mainly studied by fluorescence microscopy in single cells by overexpression of epitope-tagged proteins, due to the difficulty of detecting endogenous TLS pols, and in particular PCNAmUb. Recently, we have been able to develop specific procedures that have allowed the detection of chromatin-bound PCNAmUb and endogenous TLS pols by immunofluorescence and flow cytometry in mammalian cells (Egger, Aze et Maiorano. Cell Reports Methods 2023, see “Resources” tab). Using these procedures, we have started to study translesiosomes dynamics in single cells exposed to diverse DNA damaging agents in relation to the cell cycle stage, and to study in detail functional interactions with the DNA synthesis machinery (the replisome) at stalled DNA replication forks.
Characterization of the molecular mechanism(s) responsible for the genetic instability of very early embryos
Waqar Ali
Very early embryogenesis is surprisingly characterized by a high level of DNA damage, genomic instability and the occurrence of mutations of unclear origins. These features, which are shared with cancer cells, have been attributed to a highly contracted cell cycle, DDR inefficiency, and very recently to changes in the DNA replication timing. However, the mechanism(s) generating mutations was until recently unknown. We have tackled this issue during early Xenopus embryogenesis and shown that this depends upon robust TLS activity. Further, we have shown that both the mismatch repair pathway and, unexpectedly, an error-free homology-directed repair activity of Rad18, actively reduce the mutation load introduced by the TLS pols (Lo Furno et al., Nucleic Acids Research, 2022). We are currently exploring the contribution of TLS and other processes to the mutagenesis and genomic instability of mammalian pluripotent stem cells.


















- TopBP1 biomolecular condensates as a new therapeutic target in advanced-stage colorectal cancer. Morano L, Vezzio-Vié N, Aissanou A, Egger T, Aze A, Fiachetti S, Bordignon B, Hassen-Khodja C, Seitz H, Milazzo LA, Garambois V, Chaloin L, Bonnefoy N, Gongora C, Constantinou A, Basbous J. Elife. 2025.14:RP106196. PMID: 41117221
- Younas M, Gimenez S, Lin YL, Mettling C, Maiorano D, Reynes J, Pasero P, Rondard P, Psomas CK, Corbeau P. γ-Aminobutyric Acid-Induced Monocytic Reactive Oxygen Species Impair CD4 Restoration in Treated Adults With HIV-1. J Infect Dis. 2025, 231(5):1246-1257. PMID: 39903648.
- Coquel F, Ho SZ, Tsai KC, Yang CY, Aze A, Devin J, Chang TH, Kong-Hap M, Bioteau A, Moreaux J, Maiorano D, Pourquier P, Yang WC, Lin YL, Pasero P. Synergistic effect of inhibiting CHK2 and DNA replication on cancer cell growth. Elife. 2025, 13:RP104718. PMID: 39887032;
- Aze A, Hutchins JRA, Maiorano D. Studying Translesion DNA Synthesis Using Xenopus In Vitro Systems. Methods Mol Biol. 2024, 2740:21-36. PMID: 38393467.
- Egger T, Aze A, Maiorano D. Detection of endogenous translesion DNA synthesis in single mammalian cells. Cell Rep Methods. 2023, 3(6):100501. PMID: 37426760.
- Egger T, Aze A, Maiorano D. Protocol to analyze endogenous translesion DNA synthesis in single mammalian cells. STAR Protoc. 202, 4(3):102361. PMID: 37355992;
- Lo Furno E, Busseau I, Aze A, Lorenzi C, Saghira C, Danzi MC, Zuchner S, Maiorano D. Translesion DNA synthesis-driven mutagenesis in very early embryogenesis of fast cleaving embryos. Nucleic Acids Res. 2022, 50(2):885-898. PMID: 34939656; CNRS press release: https://www.insb.cnrs.fr/fr/cnrsinfo/un-nouveau-mecanisme-de-variabilite-genetique-durant-lembryogenese-precoce
- Lo Furno E, Recolin B, van der Laan S, Aze A, Maiorano D. Studying the DNA damage response in embryonic systems. Methods Enzymol. 2021, 661:95-120.PMID: 34776225.
- Maiorano D, El Etri J, Franchet C, Hoffmann JS. Translesion Synthesis or Repair by Specialized DNA Polymerases Limits Excessive Genomic Instability upon Replication Stress. Int J Mol Sci. 2021, 22(8):3924. PMID: 33920223.
- Basbous J, Aze A, Chaloin L, Lebdy R, Hodroj D, Ribeyre C, Larroque M, Shepard C, Kim B, Pruvost A, Moreaux J, Maiorano D, Mechali M, Constantinou A. Dihydropyrimidinase protects from DNA replication stress caused by cytotoxic metabolites. Nucleic Acids Res. 2020, 48(4):1886-1904. PMID: 31853544.
- Kermi C, Aze A, Maiorano D. Preserving Genome Integrity During the Early Embryonic DNA Replication Cycles. Genes (Basel). 2019, 10(5):398. PMID: 31137726.
- Aze A, Maiorano D. Recent advances in understanding DNA replication: cell type-specific adaptation of the DNA replication program. F1000Res. 2018, 7:F1000 Faculty Rev-1351. PMID: 30228862
- Hodroj D, Recolin B, Serhal K, Martinez S, Tsanov N, Abou Merhi R, Maiorano D. An ATR-dependent function for the Ddx19 RNA helicase in nuclear R-loop metabolism. EMBO J. 2017, 36(9):1182-1198. PMID: 28314779;
- Kermi C, Lo Furno E, Maiorano D. Regulation of DNA Replication in Early Embryonic Cleavages. Genes (Basel). 2017, 8(1):42. PMID: 28106858.
- Benkafadar N, Menardo J, Bourien J, Nouvian R, François F, Decaudin D, Maiorano D, Puel JL, Wang J. Reversible p53 inhibition prevents cisplatin ototoxicity without blocking chemotherapeutic efficacy. EMBO Mol Med. 2017, (1):7-26. PMID: 27794029. PMC5210089. Commented by Nonnekens J, Hoeijmakers JH. EMBO Mol Med. 2017 Jan;9(1):4-6. doi: 10.15252/emmm.201607062. PMID: 27852619
- Hutchins JR. Genomic Database Searching. Methods Mol Biol. 2017,1525:225-269. PMID: 27896724
- Kermi C, Prieto S, van der Laan S, Tsanov N, Recolin B, Uro-Coste E, Delisle MB, Maiorano D. RAD18 Is a Maternal Limiting Factor Silencing the UV-Dependent DNA Damage Checkpoint in Xenopus Embryos. Dev Cell. 2015, 34(3):364-72. PMID: 26212134.
- Van der Laan S, Golfetto E, Vanacker JM, Maiorano D. Cell cycle-dependent expression of Dub3, Nanog and the p160 family of nuclear receptor coactivators (NCoAs) in mouse embryonic stem cells. PLoS One. 2014, 9(4):e93663. PMID: 24695638.
- Tsanov N, Kermi C, Coulombe P, Van der Laan S, Hodroj D, Maiorano D. PIP degron proteins, substrates of CRL4Cdt2, and not PIP boxes, interfere with DNA polymerase η and κ focus formation on UV damage. Nucleic Acids Res. 2014, (6):3692-706. PMID: 24423875;
- Van der Laan S, Tsanov N, Crozet C, Maiorano D. High Dub3 expression in mouse ESCs couples the G1/S checkpoint to pluripotency. Mol Cell. 2013, 52(3):366-79. PMID: 24207026.
- Recolin B, Van der Laan S, Maiorano D. Role of replication protein A as sensor in activation of the S-phase checkpoint in Xenopus egg extracts. Nucleic Acids Res. 2012, 40(8):3431-42.PMID: 22187152.





