Research teams
Genome dynamics department
Meiosis and recombination
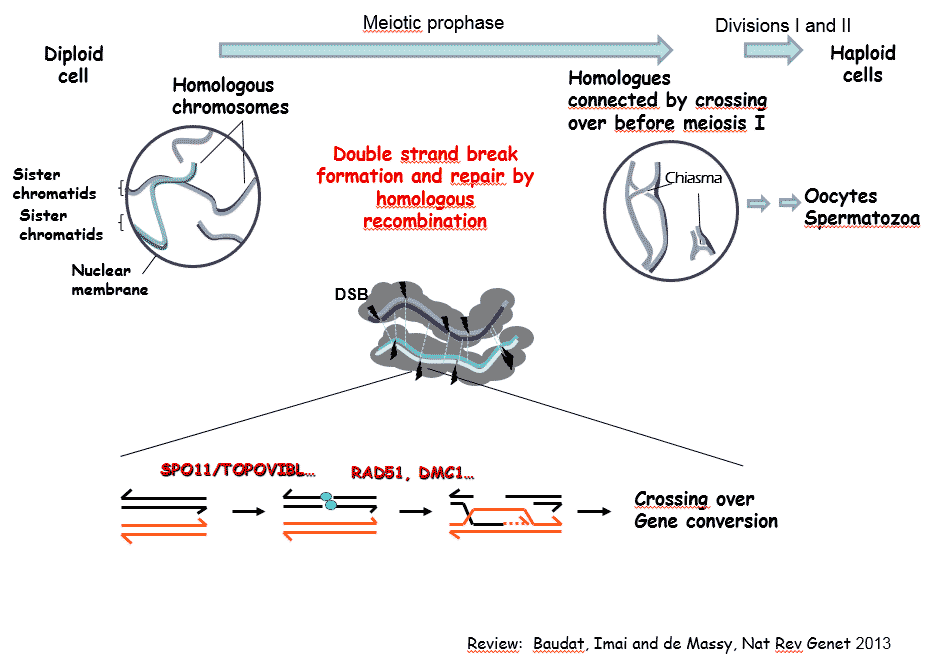
In sexually reproducing species, meiosis allows the formation of haploid gametes from diploid cells. The halving of the DNA content results from a specialized cell cycle, where a single phase of DNA replication is followed by two divisions. The reductional segregation of homologous chromosomes (homologues) at the first meiotic division requires the establishment of connections between homologues. In most species, these connections are established during a long and specialized prophase by reciprocal exchanges between homologues. These exchanges, also called crossing over, result from a highly regulated homologous recombination pathway that drives the recognition and interaction between homologues and the formation of at least one crossing over per homologue pair. Crossovers also generate new allele combinations and thus increase genetic diversity and contributes to genome evolution. The absence of crossover leads to chromosome segregation defects and sterility, and alteration of the meiotic recombination pathway can lead to genome rearrangements and aneuploidy.
Our team is investigating several aspects of the mechanism and regulation of meiotic recombination and its evolutionary implication using the mouse as a model system. Meiotic recombination events are initiated by the formation of DNA double-strand breaks (DSBs, several hundred per nucleus in mice), the repair of which leads to both crossovers and non-crossovers (gene conversion without crossover). The main steps and factors involved in this pathway are evolutionary conserved.
Bernard de Massy
ERC DSBSunrise 883605
https://cordis.europa.eu/project/id/883605
Early steps of meiotic recombination and chromosome dynamics
Corinne Grey
ANR Shape-n-Brake, ARC Breaking good)
Mechanism and control of inter-homolog recombination in meiosis
Frédéric Baudat
ANR DoRAD
























- Nosaka Y, Nagano M, Yabuta Y, Nakakita B, Nagaoka SI, Okamoto I, Sasada H, Mizuta K, Umemura F, Kuma H, Okochi Y, Katou Y, de Massy B, Inoue A, Horie A, Mandai M, Ohta H, and Saitou M (2025). Generation of germinal-vesicle oocytes from mouse embryonic stem cells under an ovarian soma-free condition. Dev Cel 10.1016/j.devcel.2025.06.008.
- Raynaud M, Sanna P, Joseph J, Clément J, Imai Y, Lareyre JJ, Laurent A, Galtier N, Baudat F, Duret L, Gagnaire PA, and de Massy B (2025). PRDM9 drives the location and rapid evolution of recombination hotspots in salmonid fish. PLoS Biol 23, e3002950. 10.1371/journal.pbio.3002950.
- Biot M, Toth A, Brun C, Guichard L, de Massy B*, and Grey C*(2024). Principles of chromosome organization for meiotic recombination. Mol Cell 84, 1826-1841 e1825. 10.1016/j.molcel.2024.04.001. INSB communication, July 2024: https://www.insb.cnrs.fr/fr/cnrsinfo/organisation-des-chromosomes-en-meiose
- Diagouraga B, Tambones I, Carivenc C, Bechara C, Nadal M, de Massy B, le Maire A C, and Robert T C (2024). The TOPOVIBL meiotic DSB formation protein: new insights from its biochemical and structural characterization. Nucleic Acids Res. 10.1093/nar/gkae587.
- Zainu A, Dupaigne P, Bouchouika S, Cau J, Clement JAJ, Auffret P, Ropars V, Charbonnier JB, de Massy B, Mercier R, Kumar R, and Baudat F C (2024). FIGNL1-FIRRM is essential for meiotic recombination and prevents DNA damage-independent RAD51 and DMC1 loading. Nat Commun 15, 7015. 10.1038/s41467-024-51458-8.
- Auffret P, de Massy B, and Clement JAJ (2024). Mapping Meiotic DNA Breaks: Two Fully-Automated Pipelines to Analyze Single-Strand DNA Sequencing Data, hotSSDS and hotSSDS-extra. Methods Mol Biol 2770, 227-261. 10.1007/978-1-0716-3698-5_16.
- Laroussi H, Juarez-Martinez AB, Le Roy A, Boeri Erba E, Gabel F, de Massy B, and Kadlec J C (2023). Characterization of the REC114-MEI4-IHO1 complex regulating meiotic DNA double-strand break formation. EMBO J, e113866. 10.15252/embj.2023113866. CNRS-Blog Focus Science, March 2025: https://lejournal.cnrs.fr/nos-blogs/focus-sciences/eclairer-les-phases-cachees-de-la-meiose
- Brinkmeier J, Coelho S, de Massy B C, and Bourbon HM (2022). Evolution and Diversity of the TopoVI and TopoVI-like Subunits With Extensive Divergence of the TOPOVIBL subunit. Mol Biol Evol 10.1093/molbev/msac227. INSB communication, February 2023: https://www.insb.cnrs.fr/fr/cnrsinfo/evolution-et-fonction-de-ciseaux-moleculaires-les-topoisomerases-et-la-meiose
- Nore A, Juarez-Martinez AB, Clement J, Brun C, Diagouraga B, Laroussi H, Grey C, Bourbon HM, Kadlec J C, Robert T C, and de Massy B C (2022). TOPOVIBL-REC114 interaction regulates meiotic DNA double-strand breaks. Nat Commun 13, 7048. 10.1038/s41467-022-34799-0.
- Dereli I*, Stanzione M*, Olmeda F, Papanikos F, Baumann M, Demir S, Carofiglio F, Lange J, de Massy B, Baarends WM, Turner J, Rulands S, and Toth A C (2021). Four-pronged negative feedback of DSB machinery in meiotic DNA-break control in mice. Nucleic Acids Res 49, 2609-2628. 10.1093/nar/gkab082.
- Mihola O, Landa V, Pratto F, Brick K, Kobets T, Kusari F, Gasic S, Smagulova F, Grey C, Flachs P, Gergelits V, Tresnak K, Silhavy J, Mlejnek P, Camerini-Otero RD, Pravenec M, Petukhova GV, and Trachtulec Z C (2021). Rat PRDM9 shapes recombination landscapes, duration of meiosis, gametogenesis, and age of fertility. BMC Biol 19, 86. 10.1186/s12915-021-01017-0.
- Imai Y*, Biot M*, Clément J, Teragaki M, Urbach S, Robert T, Baudat F, Grey C*, and de Massy B* (2020). PRDM9 activity depends on HELLS and promotes local 5-hydroxymethylcytosine enrichment. eLife 10.7554/eLife.57117.
- Papanikos F*, Clement JAJ*, Testa E, Ravindranathan R, Grey C, Dereli I, Bondarieva A, Valerio-Cabrera S, Stanzione M, Schleiffer A, Jansa P, Lustyk D, Fei JF, Adams IR, Forejt J, Barchi M, de Massy B C, and Toth A C (2019). Mouse ANKRD31 Regulates Spatiotemporal Patterning of Meiotic Recombination Initiation and Ensures Recombination between X and Y Sex Chromosomes. Mol Cell 74, 1069-1085 e1011. 10.1016/j.molcel.2019.03.022.
- Baudat F C, de Massy B, and Veyrunes F C (2019). Sex chromosome quadrivalents in oocytes of the African pygmy mouse Mus minutoides that harbors non-conventional sex chromosomes. Chromosoma 128, 397-411. 10.1007/s00412-019-00699-4.







